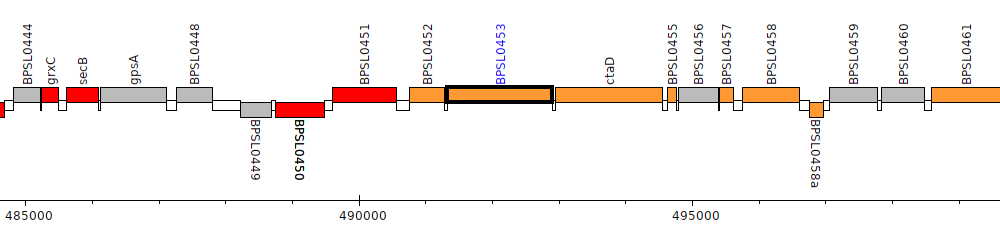
Burkholderia pseudomallei K96243, BPSL0453
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF00116
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF46626
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PS50999
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009279 | cell outer membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PR01021
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005507 | copper ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00116
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0020037 | heme binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF46626
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004129 | cytochrome-c oxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00116
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02866
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0022900 | electron transport chain |
Inferred from Sequence Model
Term mapped from: InterPro:PS50999
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00190 | Oxidative phosphorylation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF02790 | Cytochrome C oxidase subunit II, transmembrane domain | IPR011759 | Cytochrome C oxidase subunit II, transmembrane domain | 44 | 124 | 9.1E-17 |
| PRINTS | PR01166 | Cytochrome c oxidase subunit II signature | 247 | 264 | 1.3E-35 | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 375 | 408 | - | ||
| Pfam | PF00116 | Cytochrome C oxidase subunit II, periplasmic domain | IPR002429 | Cytochrome c oxidase subunit II-like C-terminal | 139 | 264 | 4.4E-33 |
| ProSiteProfiles | PS51123 | OmpA-like domain profile. | IPR006665 | OmpA-like domain | 416 | 525 | 27.135 |
| PRINTS | PR01021 | OMPA domain signature | IPR006664 | Outer membrane protein, bacterial | 429 | 451 | 3.9E-7 |
| Gene3D | G3DSA:1.10.760.10 | IPR036909 | Cytochrome c-like domain superfamily | 279 | 400 | 9.2E-26 | |
| CDD | cd07185 | OmpA_C-like | IPR006665 | OmpA-like domain | 427 | 521 | 6.19189E-20 |
| SUPERFAMILY | SSF103088 | IPR036737 | OmpA-like domain superfamily | 422 | 521 | 6.67E-20 | |
| PRINTS | PR01166 | Cytochrome c oxidase subunit II signature | 209 | 229 | 1.3E-35 | ||
| PRINTS | PR01166 | Cytochrome c oxidase subunit II signature | 185 | 206 | 1.3E-35 | ||
| SUPERFAMILY | SSF46626 | IPR036909 | Cytochrome c-like domain superfamily | 276 | 380 | 8.73E-22 | |
| SUPERFAMILY | SSF81464 | IPR036257 | Cytochrome C oxidase subunit II, transmembrane domain superfamily | 40 | 133 | 5.23E-28 | |
| Pfam | PF00691 | OmpA family | IPR006665 | OmpA-like domain | 428 | 501 | 3.0E-15 |
| SUPERFAMILY | SSF49503 | IPR008972 | Cupredoxin | 131 | 321 | 2.13E-48 | |
| TIGRFAM | TIGR02866 | CoxB: cytochrome c oxidase, subunit II | IPR014222 | Cytochrome c oxidase, subunit II | 52 | 274 | 5.4E-57 |
| ProSiteProfiles | PS51007 | Cytochrome c family profile. | IPR009056 | Cytochrome c-like domain | 296 | 372 | 9.363 |
| Gene3D | G3DSA:2.60.40.420 | IPR008972 | Cupredoxin | 135 | 274 | 9.6E-78 | |
| ProSiteProfiles | PS50857 | Cytochrome oxidase subunit II copper A binding domain profile. | IPR002429 | Cytochrome c oxidase subunit II-like C-terminal | 136 | 276 | 36.863 |
| PRINTS | PR01166 | Cytochrome c oxidase subunit II signature | 113 | 133 | 1.3E-35 | ||
| PRINTS | PR01166 | Cytochrome c oxidase subunit II signature | 101 | 113 | 1.3E-35 | ||
| ProSiteProfiles | PS50999 | Cytochrome oxidase subunit II transmembrane region profile. | IPR011759 | Cytochrome C oxidase subunit II, transmembrane domain | 40 | 135 | 17.524 |
| PRINTS | PR01021 | OMPA domain signature | IPR006664 | Outer membrane protein, bacterial | 459 | 474 | 3.9E-7 |
| Pfam | PF13442 | Cytochrome C oxidase, cbb3-type, subunit III | IPR009056 | Cytochrome c-like domain | 297 | 367 | 5.4E-11 |
| Gene3D | G3DSA:3.30.1330.60 | IPR036737 | OmpA-like domain superfamily | 401 | 521 | 2.4E-20 | |
| PRINTS | PR01166 | Cytochrome c oxidase subunit II signature | 135 | 154 | 1.3E-35 | ||
| ProSitePatterns | PS00078 | CO II and nitrous oxide reductase dinuclear copper centers signature. | IPR001505 | Copper centre Cu(A) | 210 | 258 | - |
| Gene3D | G3DSA:1.10.287.90 | IPR036257 | Cytochrome C oxidase subunit II, transmembrane domain superfamily | 63 | 134 | 9.6E-78 | |
| PRINTS | PR01021 | OMPA domain signature | IPR006664 | Outer membrane protein, bacterial | 474 | 490 | 3.9E-7 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.