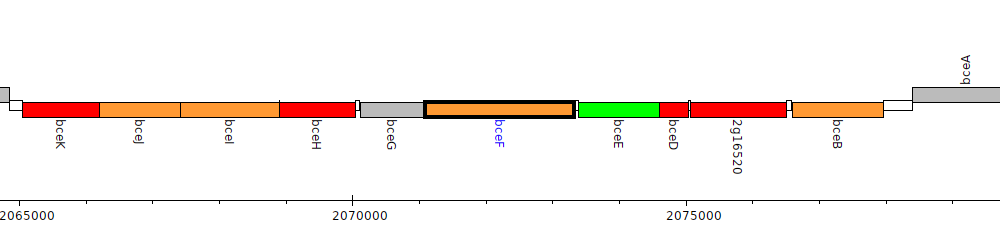
Burkholderia glumae BGR1, bglu_2g16490 (bceF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004713 | protein tyrosine kinase activity |
Inferred from Sequence Orthology
Term mapped from: GenBank:ABD78148.1
|
ECO:0000266 sequence orthology evidence used in manual assertion |
17114319 | Reviewed by curator |
| Biological Process | GO:0009103 | lipopolysaccharide biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02706
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0045226 | extracellular polysaccharide biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01007
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF02706
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Coils | Coil | 365 | 385 | - | |||
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 549 | 732 | 2.43E-37 | |
| TIGRFAM | TIGR01007 | eps_fam: capsular exopolysaccharide family | IPR005702 | Exopolysaccharide synthesis protein | 530 | 729 | 5.9E-42 |
| Pfam | PF13807 | G-rich domain on putative tyrosine kinase | IPR032807 | Tyrosine kinase, G-rich domain | 392 | 471 | 3.8E-25 |
| Coils | Coil | 337 | 357 | - | |||
| CDD | cd05387 | BY-kinase | 530 | 719 | 1.76649E-68 | ||
| Pfam | PF13614 | AAA domain | IPR025669 | AAA domain | 559 | 672 | 6.5E-11 |
| Gene3D | G3DSA:3.40.50.300 | 466 | 739 | 3.2E-71 | |||
| Pfam | PF02706 | Chain length determinant protein | IPR003856 | Polysaccharide chain length determinant N-terminal domain | 23 | 114 | 4.5E-12 |
| Coils | Coil | 293 | 313 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.