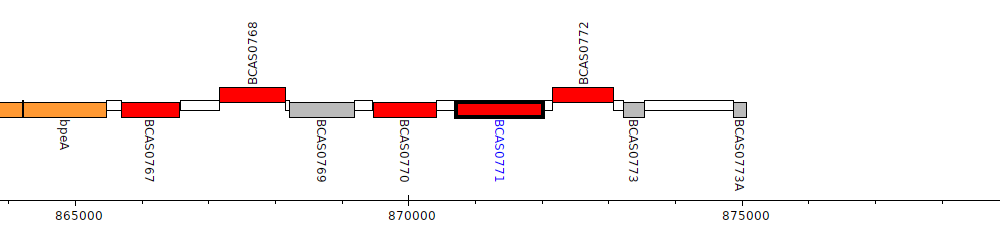
Burkholderia cenocepacia J2315, BCAS0771
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005525 | GTP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS01266
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004019 | adenylosuccinate synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00011
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006164 | purine nucleotide biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00011
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00250 | Alanine, aspartate and glutamate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | adenosine ribonucleotides <i>de novo</i> biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00230 | Purine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00250 | Alanine, aspartate and glutamate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.440.10 | IPR042109 | Adenylosuccinate synthetase, domain 1 | 5 | 255 | 1.3E-85 | |
| ProSitePatterns | PS00513 | Adenylosuccinate synthetase active site. | IPR033128 | Adenylosuccinate synthase, active site | 131 | 142 | - |
| Gene3D | G3DSA:1.10.300.10 | IPR042110 | Adenylosuccinate synthetase, domain 2 | 101 | 198 | 1.3E-85 | |
| ProSitePatterns | PS01266 | Adenylosuccinate synthetase GTP-binding site. | IPR018220 | Adenylosuccinate synthase, GTP-binding site | 10 | 17 | - |
| TIGRFAM | TIGR00184 | purA: adenylosuccinate synthase | IPR001114 | Adenylosuccinate synthetase | 5 | 426 | 7.2E-146 |
| SMART | SM00788 | IPR001114 | Adenylosuccinate synthetase | 3 | 421 | 3.6E-254 | |
| Pfam | PF00709 | Adenylosuccinate synthetase | IPR001114 | Adenylosuccinate synthetase | 4 | 420 | 8.9E-160 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 1 | 427 | 5.52E-155 | |
| CDD | cd03108 | AdSS | IPR001114 | Adenylosuccinate synthetase | 2 | 421 | 1.43426E-137 |
| Gene3D | G3DSA:3.90.170.10 | IPR042111 | Adenylosuccinate synthetase, domain 3 | 264 | 428 | 9.1E-64 | |
| Hamap | MF_00011 | Adenylosuccinate synthetase [purA]. | IPR001114 | Adenylosuccinate synthetase | 1 | 432 | 187.5 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.