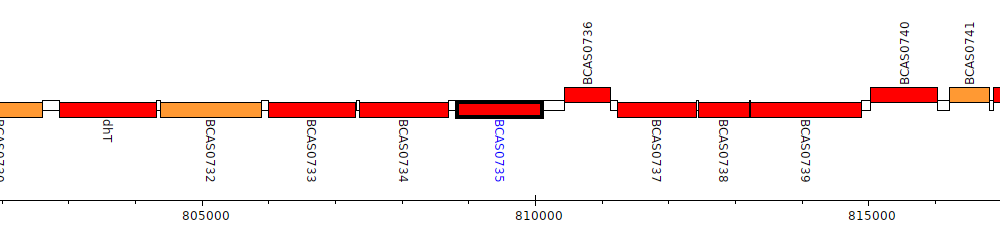
Burkholderia cenocepacia J2315, BCAS0735
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF001235
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01546
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF53187 | 15 | 419 | 4.29E-94 | |||
| CDD | cd03884 | M20_bAS | 24 | 418 | 0.0 | ||
| SUPERFAMILY | SSF55031 | IPR036264 | Bacterial exopeptidase dimerisation domain | 225 | 341 | 2.84E-30 | |
| TIGRFAM | TIGR01879 | hydantase: amidase, hydantoinase/carbamoylase family | IPR010158 | Amidase, carbamoylase-type | 22 | 418 | 3.2E-121 |
| Gene3D | G3DSA:3.30.70.360 | 225 | 339 | 9.0E-160 | |||
| Pfam | PF01546 | Peptidase family M20/M25/M40 | IPR002933 | Peptidase M20 | 93 | 417 | 3.0E-38 |
| Gene3D | G3DSA:3.40.630.10 | 31 | 415 | 9.0E-160 | |||
| PIRSF | PIRSF001235 | IPR010158 | Amidase, carbamoylase-type | 16 | 425 | 3.8E-131 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.