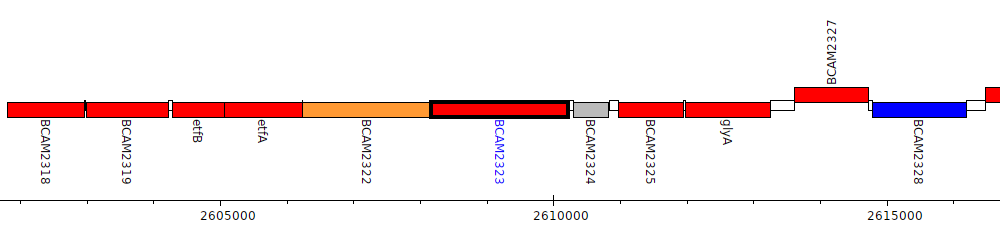
Burkholderia cenocepacia J2315, BCAM2323
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00724
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.20.20.70
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00724
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00724
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0010181 | FMN binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00724
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00724
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0010181 | FMN binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00724
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.20.20.70
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 661 | 668 | 5.6E-6 | ||
| SUPERFAMILY | SSF51395 | 1 | 374 | 1.76E-101 | |||
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 608 | 622 | 5.6E-6 | ||
| SUPERFAMILY | SSF51905 | IPR036188 | FAD/NAD(P)-binding domain superfamily | 367 | 684 | 3.5E-29 | |
| Gene3D | G3DSA:3.40.50.720 | 391 | 630 | 1.9E-61 | |||
| Gene3D | G3DSA:3.20.20.70 | IPR013785 | Aldolase-type TIM barrel | 1 | 384 | 9.8E-111 | |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | 392 | 411 | 2.6E-7 | ||
| CDD | cd04734 | OYE_like_3_FMN | 6 | 352 | 3.58927E-171 | ||
| Gene3D | G3DSA:3.50.50.60 | IPR036188 | FAD/NAD(P)-binding domain superfamily | 486 | 615 | 1.9E-61 | |
| Pfam | PF00724 | NADH:flavin oxidoreductase / NADH oxidase family | IPR001155 | NADH:flavin oxidoreductase/NADH oxidase, N-terminal | 6 | 344 | 1.1E-61 |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 391 | 413 | 5.6E-6 | ||
| Pfam | PF07992 | Pyridine nucleotide-disulphide oxidoreductase | IPR023753 | FAD/NAD(P)-binding domain | 391 | 636 | 2.7E-11 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | 607 | 623 | 2.6E-7 | ||
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | 474 | 492 | 2.6E-7 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.