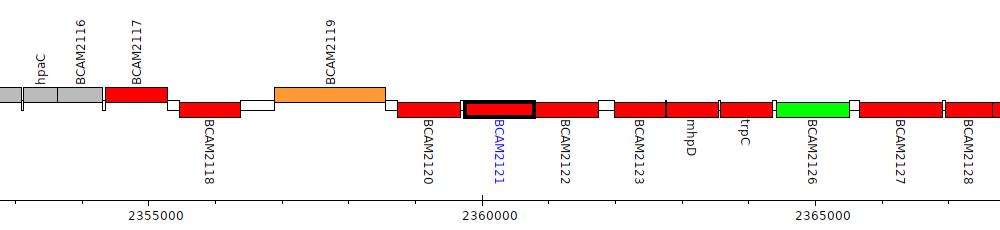
Burkholderia cenocepacia J2315, BCAM2121
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.20.20.70
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016833 | oxo-acid-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07836
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008701 | 4-hydroxy-2-oxovalerate aldolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01656
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019439 | aromatic compound catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF07836
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00621 | Dioxin degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | 2-hydroxypenta-2,4-dienoate degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00360 | Phenylalanine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00362 | Benzoate degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01220 | Degradation of aromatic compounds | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00622 | Xylene degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00621 | Dioxin degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00622 | Xylene degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00362 | Benzoate degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00360 | Phenylalanine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF07836 | DmpG-like communication domain | IPR012425 | DmpG-like communication | 275 | 336 | 9.5E-30 |
| Hamap | MF_01656 | 4-hydroxy-2-oxovalerate aldolase [mhpE]. | IPR017629 | 4-hydroxy-2-oxovalerate aldolase | 1 | 344 | 220.767 |
| SUPERFAMILY | SSF51569 | 3 | 278 | 8.06E-68 | |||
| Pfam | PF00682 | HMGL-like | IPR000891 | Pyruvate carboxyltransferase | 8 | 264 | 3.7E-74 |
| TIGRFAM | TIGR03217 | 4OH_2_O_val_ald: 4-hydroxy-2-oxovalerate aldolase | IPR017629 | 4-hydroxy-2-oxovalerate aldolase | 6 | 338 | 1.8E-177 |
| CDD | cd07943 | DRE_TIM_HOA | IPR035685 | 4-hydroxy-2-oxovalerate aldolase, N-terminal catalytic TIM barrel domain | 8 | 271 | 3.43608E-157 |
| SUPERFAMILY | SSF89000 | 290 | 339 | 1.18E-18 | |||
| ProSiteProfiles | PS50991 | Pyruvate carboxyltransferase domain. | IPR000891 | Pyruvate carboxyltransferase | 8 | 260 | 25.403 |
| Gene3D | G3DSA:3.20.20.70 | IPR013785 | Aldolase-type TIM barrel | 1 | 270 | 6.2E-90 | |
| Gene3D | G3DSA:1.10.8.60 | 277 | 342 | 2.0E-32 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.