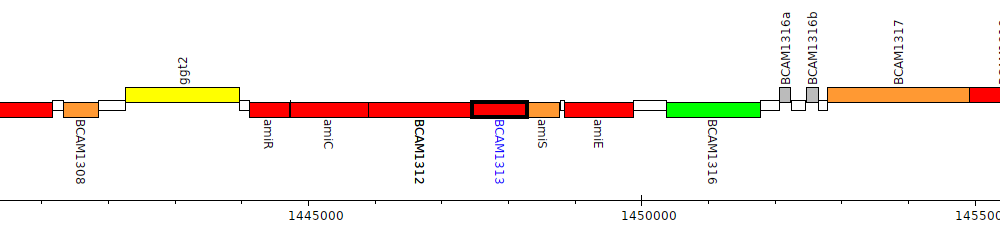
Burkholderia cenocepacia J2315, BCAM1313
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF07728
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07728
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF07728 | AAA domain (dynein-related subfamily) | IPR011704 | ATPase, dynein-related, AAA domain | 43 | 177 | 1.3E-30 |
| CDD | cd00009 | AAA | 39 | 116 | 6.32272E-5 | ||
| Gene3D | G3DSA:3.40.50.300 | 30 | 195 | 5.5E-30 | |||
| Pfam | PF08406 | CbbQ/NirQ/NorQ C-terminal | IPR013615 | CbbQ/NirQ/NorQ, C-terminal | 190 | 273 | 3.3E-28 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 36 | 241 | 1.87E-35 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.