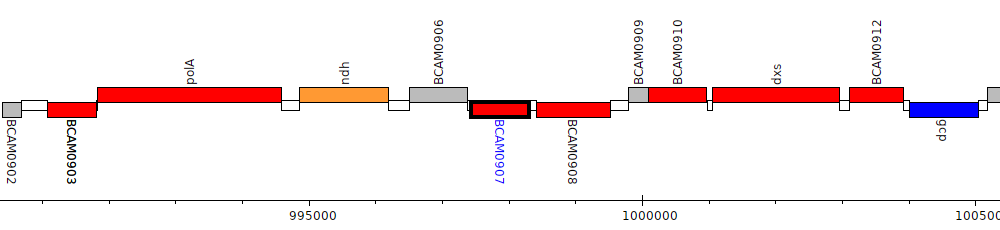
Burkholderia cenocepacia J2315, BCAM0907
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004792 | thiosulfate sulfurtransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00683
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | sulfide oxidation IV (metazoa) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00920 | Sulfur metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00270 | Cysteine and methionine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00920 | Sulfur metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj04122 | Sulfur relay system | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | thiosulfate disproportionation IV (rhodanese) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| ProSitePatterns | PS00683 | Rhodanese C-terminal signature. | IPR001307 | Thiosulphate sulfurtransferase, conserved site | 265 | 275 | - |
| SUPERFAMILY | SSF52821 | IPR036873 | Rhodanese-like domain superfamily | 166 | 288 | 1.7E-31 | |
| CDD | cd01448 | TST_Repeat_1 | 9 | 136 | 2.09179E-36 | ||
| ProSiteProfiles | PS50206 | Rhodanese domain profile. | IPR001763 | Rhodanese-like domain | 171 | 284 | 16.777 |
| ProSiteProfiles | PS50206 | Rhodanese domain profile. | IPR001763 | Rhodanese-like domain | 22 | 141 | 20.831 |
| CDD | cd01449 | TST_Repeat_2 | 161 | 277 | 1.27325E-45 | ||
| Gene3D | G3DSA:3.40.250.10 | IPR036873 | Rhodanese-like domain superfamily | 165 | 286 | 2.4E-32 | |
| Pfam | PF00581 | Rhodanese-like domain | IPR001763 | Rhodanese-like domain | 17 | 134 | 3.4E-17 |
| SMART | SM00450 | IPR001763 | Rhodanese-like domain | 161 | 281 | 8.8E-14 | |
| ProSitePatterns | PS00380 | Rhodanese signature 1. | IPR001307 | Thiosulphate sulfurtransferase, conserved site | 45 | 56 | - |
| Gene3D | G3DSA:3.40.250.10 | IPR036873 | Rhodanese-like domain superfamily | 2 | 155 | 2.4E-46 | |
| Pfam | PF00581 | Rhodanese-like domain | IPR001763 | Rhodanese-like domain | 168 | 277 | 1.8E-9 |
| SMART | SM00450 | IPR001763 | Rhodanese-like domain | 10 | 138 | 4.7E-19 | |
| SUPERFAMILY | SSF52821 | IPR036873 | Rhodanese-like domain superfamily | 2 | 153 | 6.16E-43 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.