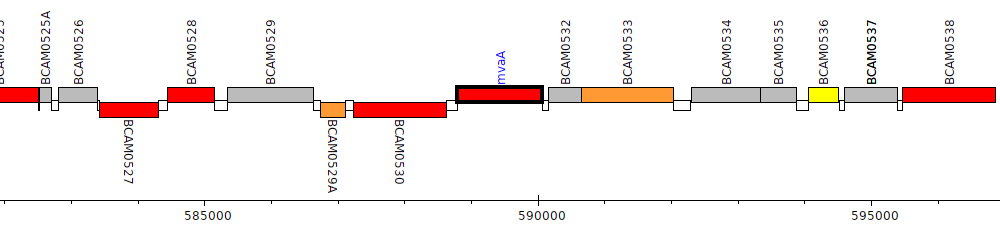
Burkholderia cenocepacia J2315, BCAM0531 (mvaA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0015936 | coenzyme A metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00532
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PS00318
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050662 | coenzyme binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00532
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00532
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004420 | hydroxymethylglutaryl-CoA reductase (NADPH) activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00318
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | mevalonate pathway III (archaea) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | isoprene biosynthesis II (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | mevalonate pathway I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | mevalonate pathway II (archaea) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00900 | Terpenoid backbone biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00900 | Terpenoid backbone biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.30.70.420 | IPR009023 | Hydroxymethylglutaryl-CoA reductase, class I/II, NAD/NADP-binding domain superfamily | 110 | 220 | 4.6E-134 | |
| Gene3D | G3DSA:3.90.770.10 | IPR023074 | Hydroxymethylglutaryl-CoA reductase, class I/II, catalytic domain superfamily | 15 | 372 | 4.6E-134 | |
| PRINTS | PR00071 | Hydroxymethylglutaryl-coenzyme A reductase signature | IPR002202 | Hydroxymethylglutaryl-CoA reductase, class I/II | 313 | 334 | 1.4E-5 |
| ProSiteProfiles | PS50065 | Hydroxymethylglutaryl-coenzyme A reductases family profile. | IPR002202 | Hydroxymethylglutaryl-CoA reductase, class I/II | 3 | 385 | 39.854 |
| SUPERFAMILY | SSF56542 | IPR009029 | Hydroxymethylglutaryl-CoA reductase, class I/II, substrate-binding domain superfamily | 7 | 423 | 6.67E-102 | |
| PRINTS | PR00071 | Hydroxymethylglutaryl-coenzyme A reductase signature | IPR002202 | Hydroxymethylglutaryl-CoA reductase, class I/II | 77 | 97 | 1.4E-5 |
| ProSitePatterns | PS00318 | Hydroxymethylglutaryl-coenzyme A reductases signature 2. | IPR023076 | Hydroxymethylglutaryl-CoA reductase, class I/II, conserved site | 325 | 332 | - |
| TIGRFAM | TIGR00532 | HMG_CoA_R_NAD: hydroxymethylglutaryl-CoA reductase, degradative | IPR004553 | Hydroxymethylglutaryl-CoA reductase, bacterial-type | 6 | 386 | 4.6E-123 |
| ProSitePatterns | PS00066 | Hydroxymethylglutaryl-coenzyme A reductases signature 1. | IPR023076 | Hydroxymethylglutaryl-CoA reductase, class I/II, conserved site | 176 | 190 | - |
| CDD | cd00644 | HMG-CoA_reductase_classII | IPR004553 | Hydroxymethylglutaryl-CoA reductase, bacterial-type | 8 | 423 | 0.0 |
| SUPERFAMILY | SSF55035 | IPR009023 | Hydroxymethylglutaryl-CoA reductase, class I/II, NAD/NADP-binding domain superfamily | 111 | 219 | 7.85E-32 | |
| PRINTS | PR00071 | Hydroxymethylglutaryl-coenzyme A reductase signature | IPR002202 | Hydroxymethylglutaryl-CoA reductase, class I/II | 50 | 71 | 1.4E-5 |
| ProSitePatterns | PS01192 | Hydroxymethylglutaryl-coenzyme A reductases signature 3. | IPR023076 | Hydroxymethylglutaryl-CoA reductase, class I/II, conserved site | 371 | 384 | - |
| PRINTS | PR00071 | Hydroxymethylglutaryl-coenzyme A reductase signature | IPR002202 | Hydroxymethylglutaryl-CoA reductase, class I/II | 174 | 192 | 1.4E-5 |
| Pfam | PF00368 | Hydroxymethylglutaryl-coenzyme A reductase | IPR002202 | Hydroxymethylglutaryl-CoA reductase, class I/II | 16 | 386 | 2.5E-109 |
| Gene3D | G3DSA:1.10.8.660 | 376 | 425 | 2.2E-13 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.