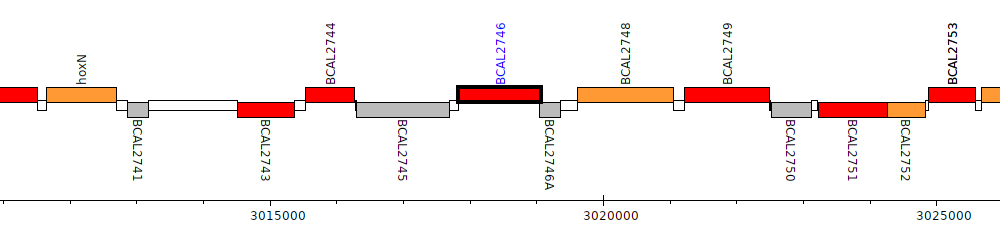
Burkholderia cenocepacia J2315, BCAL2746
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer |
Inferred from Sequence Model
Term mapped from: InterPro:PR00143
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01210 | 2-Oxocarboxylic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00020 | Citrate cycle (TCA cycle) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00630 | Glyoxylate and dicarboxylate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF46955 | IPR009061 | Putative DNA-binding domain superfamily | 4 | 53 | 5.49E-7 | |
| CDD | cd06102 | citrate_synt_like_2 | 62 | 405 | 5.98979E-95 | ||
| Gene3D | G3DSA:1.10.1660.10 | 2 | 66 | 5.3E-6 | |||
| PRINTS | PR00143 | Citrate synthase signature | IPR002020 | Citrate synthase | 149 | 162 | 1.1E-20 |
| PRINTS | PR00143 | Citrate synthase signature | IPR002020 | Citrate synthase | 350 | 366 | 1.1E-20 |
| Gene3D | G3DSA:1.10.230.10 | IPR016143 | Citrate synthase-like, small alpha subdomain | 262 | 364 | 3.3E-47 | |
| SUPERFAMILY | SSF48256 | IPR036969 | Citrate synthase superfamily | 55 | 406 | 1.7E-70 | |
| PRINTS | PR00143 | Citrate synthase signature | IPR002020 | Citrate synthase | 214 | 229 | 1.1E-20 |
| Pfam | PF00285 | Citrate synthase, C-terminal domain | IPR002020 | Citrate synthase | 70 | 400 | 9.0E-70 |
| Gene3D | G3DSA:1.10.580.10 | IPR016142 | Citrate synthase-like, large alpha subdomain | 87 | 390 | 3.3E-47 | |
| PRINTS | PR00143 | Citrate synthase signature | IPR002020 | Citrate synthase | 236 | 264 | 1.1E-20 |
| PRINTS | PR00143 | Citrate synthase signature | IPR002020 | Citrate synthase | 370 | 384 | 1.1E-20 |
| Pfam | PF12728 | Helix-turn-helix domain | IPR041657 | Helix-turn-helix domain, group 17 | 4 | 53 | 2.4E-7 |
| PRINTS | PR00143 | Citrate synthase signature | IPR002020 | Citrate synthase | 296 | 316 | 1.1E-20 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.