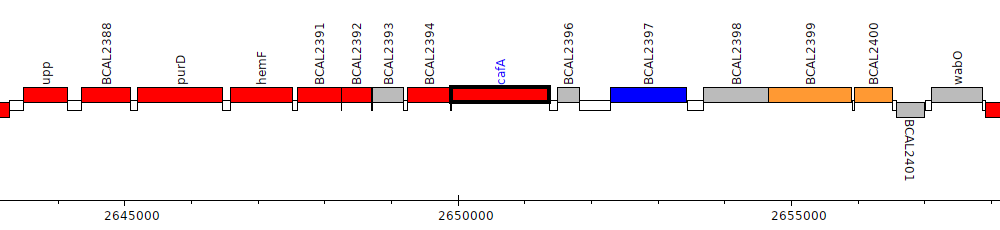
Burkholderia cenocepacia J2315, BCAL2395 (cafA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006396 | RNA processing |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00757
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004540 | ribonuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00757
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF10150
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR00757 | RNaseEG: ribonuclease, Rne/Rng family | IPR004659 | Ribonuclease E/G | 13 | 426 | 3.9E-156 |
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 39 | 126 | 4.54E-18 | |
| CDD | cd04453 | S1_RNase_E | 38 | 126 | 5.77682E-36 | ||
| Gene3D | G3DSA:2.40.50.140 | 31 | 126 | 1.1E-28 | |||
| Pfam | PF10150 | Ribonuclease E/G family | IPR019307 | RNA-binding protein AU-1/Ribonuclease E/G | 122 | 393 | 3.6E-103 |
| SMART | SM00316 | IPR022967 | RNA-binding domain, S1 | 37 | 120 | 4.8E-6 | |
| Gene3D | G3DSA:3.40.1260.20 | 402 | 488 | 1.6E-5 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.