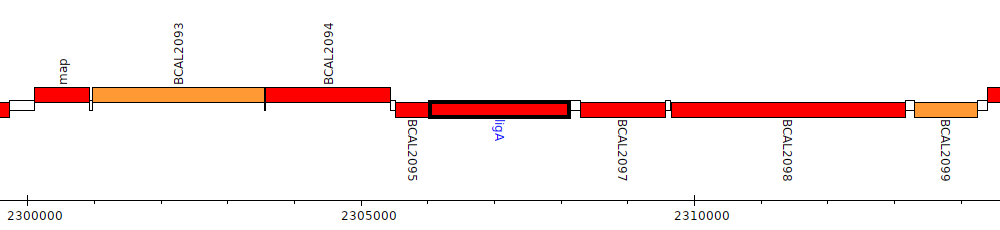
Burkholderia cenocepacia J2315, BCAL2096 (ligA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00278
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006260 | DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:PF03120
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006281 | DNA repair |
Inferred from Sequence Model
Term mapped from: InterPro:PF03120
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003911 | DNA ligase (NAD+) activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd00114
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj03430 | Mismatch repair | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03030 | DNA replication | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03420 | Nucleotide excision repair | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03410 | Base excision repair | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF12826 | Helix-hairpin-helix motif | IPR041663 | DisA/LigA, helix-hairpin-helix motif | 527 | 590 | 3.3E-27 |
| Gene3D | G3DSA:3.30.470.90 | 78 | 334 | 1.6E-95 | |||
| SUPERFAMILY | SSF52113 | IPR036420 | BRCT domain superfamily | 610 | 687 | 1.64E-20 | |
| SUPERFAMILY | SSF56091 | 13 | 333 | 3.2E-116 | |||
| PIRSF | PIRSF001604 | IPR001679 | NAD-dependent DNA ligase | 5 | 690 | 6.7E-250 | |
| Gene3D | G3DSA:1.10.150.20 | 449 | 521 | 6.4E-29 | |||
| SMART | SM00278 | IPR003583 | Helix-hairpin-helix DNA-binding motif, class 1 | 497 | 516 | 150.0 | |
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 335 | 416 | 1.71E-28 | |
| CDD | cd00114 | LIGANc | IPR013839 | NAD-dependent DNA ligase, adenylation | 15 | 334 | 4.03553E-166 |
| ProSitePatterns | PS01056 | NAD-dependent DNA ligase signature 2. | IPR033136 | NAD-dependent DNA ligase, conserved site | 348 | 363 | - |
| Pfam | PF14520 | Helix-hairpin-helix domain | 464 | 519 | 4.8E-4 | ||
| SUPERFAMILY | SSF47781 | IPR010994 | RuvA domain 2-like | 420 | 597 | 1.07E-65 | |
| Gene3D | G3DSA:2.40.50.140 | 335 | 406 | 1.4E-33 | |||
| SMART | SM00278 | IPR003583 | Helix-hairpin-helix DNA-binding motif, class 1 | 463 | 482 | 86.0 | |
| Pfam | PF03119 | NAD-dependent DNA ligase C4 zinc finger domain | IPR004149 | Zinc-finger, NAD-dependent DNA ligase C4-type | 424 | 451 | 6.8E-12 |
| ProSiteProfiles | PS50172 | BRCT domain profile. | IPR001357 | BRCT domain | 610 | 691 | 15.927 |
| Pfam | PF03120 | NAD-dependent DNA ligase OB-fold domain | IPR004150 | NAD-dependent DNA ligase, OB-fold | 339 | 415 | 1.4E-35 |
| ProSitePatterns | PS01055 | NAD-dependent DNA ligase signature 1. | IPR018239 | NAD-dependent DNA ligase, active site | 132 | 161 | - |
| Pfam | PF01653 | NAD-dependent DNA ligase adenylation domain | IPR013839 | NAD-dependent DNA ligase, adenylation | 15 | 335 | 4.0E-123 |
| SMART | SM00532 | IPR013840 | NAD-dependent DNA ligase, N-terminal | 12 | 465 | 1.0E-263 | |
| SMART | SM00292 | IPR001357 | BRCT domain | 612 | 689 | 4.6E-16 | |
| Pfam | PF00533 | BRCA1 C Terminus (BRCT) domain | IPR001357 | BRCT domain | 612 | 685 | 6.8E-14 |
| CDD | cd17748 | BRCT_DNA_ligase_like | 615 | 685 | 4.98406E-30 | ||
| TIGRFAM | TIGR00575 | dnlj: DNA ligase, NAD-dependent | IPR001679 | NAD-dependent DNA ligase | 20 | 681 | 1.3E-257 |
| Gene3D | G3DSA:2.20.70.80 | 408 | 448 | 7.6E-20 | |||
| Hamap | MF_01588 | DNA ligase [ligA]. | IPR001679 | NAD-dependent DNA ligase | 11 | 686 | 54.952 |
| SMART | SM00278 | IPR003583 | Helix-hairpin-helix DNA-binding motif, class 1 | 561 | 580 | 2.9 | |
| Gene3D | G3DSA:1.10.287.610 | 11 | 76 | 5.3E-28 | |||
| Gene3D | G3DSA:1.10.150.20 | 522 | 595 | 5.5E-30 | |||
| Gene3D | G3DSA:3.40.50.10190 | IPR036420 | BRCT domain superfamily | 596 | 688 | 3.9E-29 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.