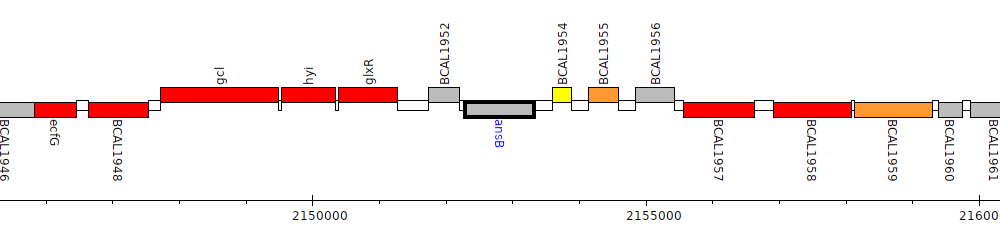
Burkholderia cenocepacia J2315, BCAL1953 (ansB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006520 | cellular amino acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PS00144
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006528 | asparagine metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd08964
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004067 | asparaginase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd08964
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00250 | Alanine, aspartate and glutamate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00250 | Alanine, aspartate and glutamate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00460 | Cyanoamino acid metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00460 | Cyanoamino acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.40 | IPR027473 | L-asparaginase, C-terminal | 226 | 339 | 6.0E-35 | |
| SFLD | SFLDS00057 | Glutaminase/Asparaginase | 13 | 335 | 0.0 | ||
| Pfam | PF17763 | Glutaminase/Asparaginase C-terminal domain | IPR040919 | Asparaginase/glutaminase, C-terminal | 227 | 337 | 1.8E-23 |
| CDD | cd08964 | L-asparaginase_II | IPR004550 | L-asparaginase, type II | 16 | 334 | 7.24165E-132 |
| SUPERFAMILY | SSF53774 | IPR036152 | Asparaginase/glutaminase-like superfamily | 14 | 338 | 1.57E-94 | |
| PRINTS | PR00139 | Asparaginase/glutaminase family signature | IPR006034 | Asparaginase/glutaminase-like | 18 | 29 | 1.7E-22 |
| PIRSF | PIRSF001220 | IPR006034 | Asparaginase/glutaminase-like | 13 | 340 | 8.0E-85 | |
| Pfam | PF00710 | Asparaginase, N-terminal | IPR027474 | L-asparaginase, N-terminal | 17 | 210 | 7.9E-61 |
| PIRSF | PIRSF500176 | 14 | 340 | 9.1E-78 | |||
| TIGRFAM | TIGR00520 | asnASE_II: L-asparaginase, type II | IPR004550 | L-asparaginase, type II | 10 | 338 | 1.4E-107 |
| PRINTS | PR00139 | Asparaginase/glutaminase family signature | IPR006034 | Asparaginase/glutaminase-like | 273 | 291 | 1.7E-22 |
| PRINTS | PR00139 | Asparaginase/glutaminase family signature | IPR006034 | Asparaginase/glutaminase-like | 98 | 116 | 1.7E-22 |
| ProSitePatterns | PS00917 | Asparaginase / glutaminase active site signature 2. | IPR027475 | Asparaginase/glutaminase, active site 2 | 99 | 109 | - |
| SMART | SM00870 | IPR006034 | Asparaginase/glutaminase-like | 17 | 333 | 4.8E-126 | |
| ProSitePatterns | PS00144 | Asparaginase / glutaminase active site signature 1. | IPR020827 | Asparaginase/glutaminase, active site 1 | 20 | 28 | - |
| Gene3D | G3DSA:3.40.50.1170 | IPR037152 | L-asparaginase, N-terminal domain superfamily | 13 | 224 | 2.5E-81 | |
| ProSiteProfiles | PS51732 | Asparaginase / glutaminase domain profile. | IPR006034 | Asparaginase/glutaminase-like | 16 | 340 | 71.115 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.