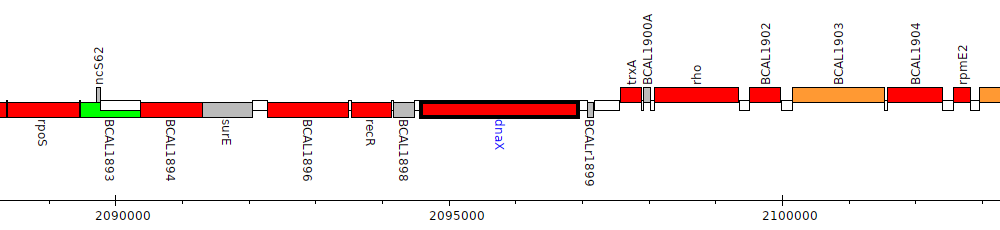
Burkholderia cenocepacia J2315, BCAL1899 (dnaX)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006260 | DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48019
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02397
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009360 | DNA polymerase III complex |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02397
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003887 | DNA-directed DNA polymerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF12169
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48019
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03440 | Homologous recombination | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03430 | Mismatch repair | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00240 | Pyrimidine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03030 | DNA replication | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 34 | 225 | 5.4E-56 | |
| TIGRFAM | TIGR02397 | dnaX_nterm: DNA polymerase III, subunit gamma and tau | IPR012763 | DNA polymerase III, subunit gamma/ tau | 3 | 355 | 5.1E-127 |
| CDD | cd00009 | AAA | 19 | 170 | 4.75922E-13 | ||
| CDD | cd18137 | HLD_clamp_pol_III_gamma_tau | 178 | 242 | 7.58027E-25 | ||
| SUPERFAMILY | SSF48019 | IPR008921 | DNA polymerase III, clamp loader complex, gamma/delta/delta subunit, C-terminal | 244 | 361 | 1.87E-31 | |
| Pfam | PF12170 | DNA polymerase III tau subunit V interacting with alpha | IPR021029 | DNA polymerase III, tau subunit, domain V | 656 | 771 | 3.8E-10 |
| Gene3D | G3DSA:3.40.50.300 | 13 | 177 | 3.2E-103 | |||
| Gene3D | G3DSA:1.10.8.60 | 8 | 239 | 3.2E-103 | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 401 | 519 | - | ||
| Gene3D | G3DSA:3.30.300.150 | IPR038249 | DNA polymerase III, tau subunit, C-terminal domain superfamily | 646 | 756 | 5.6E-8 | |
| SMART | SM00382 | IPR003593 | AAA+ ATPase domain | 37 | 179 | 5.4E-7 | |
| Pfam | PF13177 | DNA polymerase III, delta subunit | 20 | 178 | 1.1E-39 | ||
| Gene3D | G3DSA:1.20.272.10 | 242 | 362 | 3.5E-39 | |||
| Pfam | PF12169 | DNA polymerase III subunits gamma and tau domain III | IPR022754 | DNA polymerase III, gamma subunit, domain III | 232 | 358 | 2.1E-24 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 367 | 386 | - | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 533 | 589 | - | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 464 | 479 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.