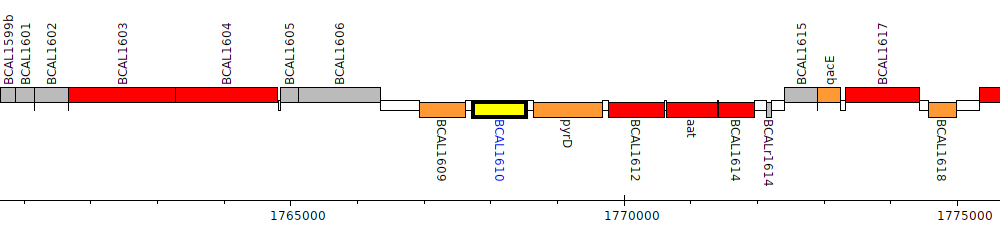
Burkholderia cenocepacia J2315, BCAL1610
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0015276 | ligand-gated ion channel activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00079
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:SM00079
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj02010 | ABC transporters | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00079 | IPR001320 | Ionotropic glutamate receptor | 38 | 257 | 0.0031 | |
| Gene3D | G3DSA:3.40.190.10 | 36 | 255 | 6.9E-71 | |||
| ProSitePatterns | PS01039 | Bacterial extracellular solute-binding proteins, family 3 signature. | IPR018313 | Solute-binding protein family 3, conserved site | 61 | 74 | - |
| SMART | SM00062 | IPR001638 | Solute-binding protein family 3/N-terminal domain of MltF | 38 | 258 | 3.9E-81 | |
| Gene3D | G3DSA:3.40.190.10 | 122 | 220 | 6.9E-71 | |||
| CDD | cd13712 | PBP2_FliY | 38 | 256 | 1.69671E-127 | ||
| Pfam | PF00497 | Bacterial extracellular solute-binding proteins, family 3 | IPR001638 | Solute-binding protein family 3/N-terminal domain of MltF | 39 | 256 | 4.0E-67 |
| SUPERFAMILY | SSF53850 | 8 | 258 | 1.37E-72 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.