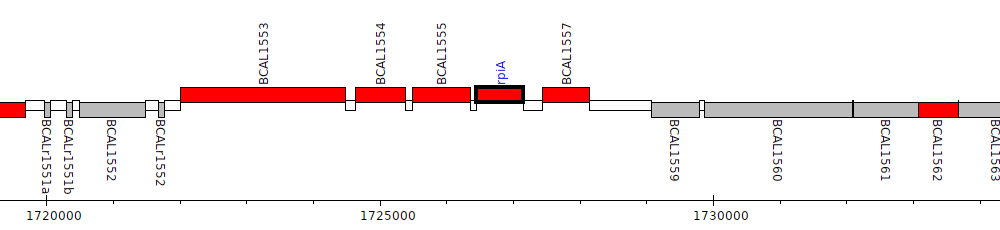
Burkholderia cenocepacia J2315, BCAL1556 (rpiA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004751 | ribose-5-phosphate isomerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00021
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009052 | pentose-phosphate shunt, non-oxidative branch |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00021
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00030 | Pentose phosphate pathway | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00051 | Fructose and mannose metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | Rubisco shunt | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00030 | Pentose phosphate pathway | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | formaldehyde assimilation II (assimilatory RuMP Cycle) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00710 | Carbon fixation in photosynthetic organisms | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00710 | Carbon fixation in photosynthetic organisms | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Hamap | MF_00170 | Ribose-5-phosphate isomerase A [rpiA]. | IPR020672 | Ribose-5-phosphate isomerase, type A, subgroup | 3 | 226 | 32.031 |
| CDD | cd01398 | RPI_A | IPR004788 | Ribose 5-phosphate isomerase, type A | 6 | 219 | 8.77426E-75 |
| Pfam | PF06026 | Ribose 5-phosphate isomerase A (phosphoriboisomerase A) | IPR004788 | Ribose 5-phosphate isomerase, type A | 53 | 222 | 1.5E-55 |
| TIGRFAM | TIGR00021 | rpiA: ribose 5-phosphate isomerase A | IPR004788 | Ribose 5-phosphate isomerase, type A | 12 | 223 | 1.5E-64 |
| Gene3D | G3DSA:3.40.50.1360 | 13 | 219 | 1.3E-75 | |||
| Gene3D | G3DSA:3.30.70.260 | 129 | 205 | 1.3E-75 | |||
| SUPERFAMILY | SSF75445 | 130 | 207 | 5.1E-21 | |||
| SUPERFAMILY | SSF100950 | IPR037171 | NagB/RpiA transferase-like | 1 | 150 | 5.71E-43 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.