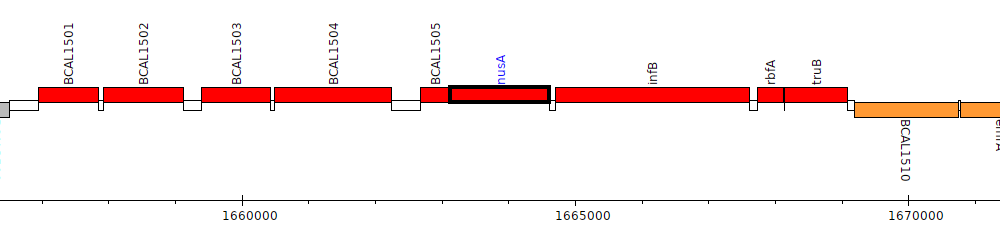
Burkholderia cenocepacia J2315, BCAL1506 (nusA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF13184
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS50126
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003700 | DNA-binding transcription factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF08529
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006353 | DNA-templated transcription, termination |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00945_B
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF47794
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0031554 | regulation of DNA-templated transcription, termination |
Inferred from Sequence Model
Term mapped from: InterPro:PF08529
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0031564 | transcription antitermination |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00945_B
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd02134 | NusA_KH | 204 | 276 | 0.00132712 | ||
| Pfam | PF00575 | S1 RNA binding domain | IPR003029 | S1 domain | 139 | 198 | 1.2E-8 |
| TIGRFAM | TIGR01954 | nusA_Cterm_rpt: transcription termination factor NusA, C-terminal duplication | IPR010214 | Transcription termination factor NusA, C-terminal duplication | 367 | 416 | 8.1E-20 |
| SUPERFAMILY | SSF47794 | IPR010995 | DNA repair Rad51/transcription factor NusA, alpha-helical | 359 | 418 | 1.02E-15 | |
| CDD | cd02134 | NusA_KH | 282 | 341 | 2.10074E-14 | ||
| SUPERFAMILY | SSF47794 | IPR010995 | DNA repair Rad51/transcription factor NusA, alpha-helical | 433 | 490 | 1.88E-12 | |
| Gene3D | G3DSA:3.30.1480.10 | IPR036555 | NusA, N-terminal domain superfamily | 1 | 132 | 4.1E-43 | |
| SUPERFAMILY | SSF54814 | IPR009019 | K homology domain superfamily, prokaryotic type | 202 | 280 | 5.69E-29 | |
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 136 | 201 | 7.17E-17 | |
| SUPERFAMILY | SSF54814 | IPR009019 | K homology domain superfamily, prokaryotic type | 281 | 344 | 2.03E-14 | |
| SUPERFAMILY | SSF69705 | IPR036555 | NusA, N-terminal domain superfamily | 1 | 129 | 5.89E-40 | |
| TIGRFAM | TIGR01953 | NusA: transcription termination factor NusA | IPR010213 | Transcription termination factor NusA | 5 | 344 | 2.9E-122 |
| SMART | SM00316 | IPR022967 | RNA-binding domain, S1 | 136 | 202 | 1.4E-9 | |
| Pfam | PF14520 | Helix-hairpin-helix domain | 432 | 487 | 5.5E-9 | ||
| ProSiteProfiles | PS50126 | S1 domain profile. | IPR003029 | S1 domain | 138 | 202 | 11.899 |
| TIGRFAM | TIGR01954 | nusA_Cterm_rpt: transcription termination factor NusA, C-terminal duplication | IPR010214 | Transcription termination factor NusA, C-terminal duplication | 443 | 489 | 1.1E-15 |
| Gene3D | G3DSA:2.40.50.140 | 137 | 200 | 2.2E-20 | |||
| CDD | cd04455 | S1_NusA | 135 | 201 | 1.6855E-23 | ||
| Hamap | MF_00945_B | Transcription termination/antitermination protein NusA [nusA]. | IPR030842 | Transcription termination/antitermination protein NusA, bacterial | 2 | 412 | 29.158 |
| Gene3D | G3DSA:1.10.150.20 | 357 | 423 | 1.5E-28 | |||
| ProSiteProfiles | PS50084 | Type-1 KH domain profile. | 304 | 340 | 9.561 | ||
| Gene3D | G3DSA:3.30.300.20 | IPR015946 | K homology domain-like, alpha/beta | 201 | 279 | 2.9E-29 | |
| Gene3D | G3DSA:3.30.300.20 | IPR015946 | K homology domain-like, alpha/beta | 280 | 347 | 6.3E-25 | |
| Coils | Coil | 405 | 425 | - | |||
| SMART | SM00322 | IPR004087 | K Homology domain | 303 | 376 | 0.0093 | |
| Gene3D | G3DSA:1.10.150.20 | 431 | 489 | 1.0E-19 | |||
| SMART | SM00322 | IPR004087 | K Homology domain | 232 | 295 | 36.0 | |
| Pfam | PF13184 | NusA-like KH domain | IPR025249 | KH domain, NusA-like | 232 | 299 | 4.1E-24 |
| Pfam | PF08529 | NusA N-terminal domain | IPR013735 | Transcription factor NusA, N-terminal | 4 | 127 | 1.3E-36 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.