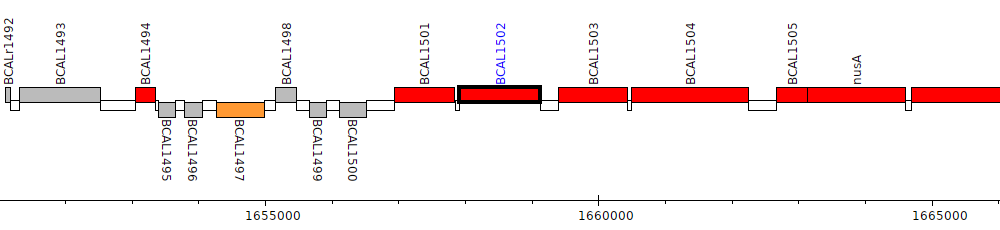
Burkholderia cenocepacia J2315, BCAL1502
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.640.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030170 | pyridoxal phosphate binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00155
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009058 | biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00155
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00155 | Aminotransferase class I and II | IPR004839 | Aminotransferase, class I/classII | 52 | 389 | 5.6E-54 |
| SUPERFAMILY | SSF53383 | IPR015424 | Pyridoxal phosphate-dependent transferase | 21 | 394 | 1.16E-105 | |
| Gene3D | G3DSA:3.40.640.10 | IPR015421 | Pyridoxal phosphate-dependent transferase, major domain | 71 | 292 | 9.6E-102 | |
| CDD | cd00609 | AAT_like | 42 | 391 | 3.43478E-97 | ||
| ProSitePatterns | PS00105 | Aminotransferases class-I pyridoxal-phosphate attachment site. | IPR004838 | Aminotransferases, class-I, pyridoxal-phosphate-binding site | 242 | 255 | - |
| Gene3D | G3DSA:3.90.1150.10 | IPR015422 | Pyridoxal phosphate-dependent transferase domain 1 | 40 | 389 | 9.6E-102 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.