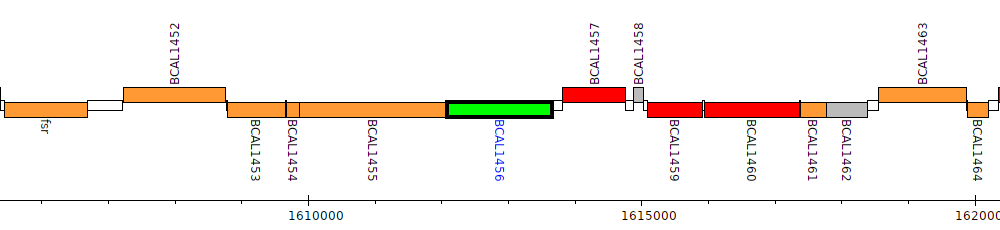
Burkholderia cenocepacia J2315, BCAL1456
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01845
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0015562 | efflux transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02321
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF02321
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01845
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Coils | Coil | 85 | 105 | - | |||
| Pfam | PF02321 | Outer membrane efflux protein | IPR003423 | Outer membrane efflux protein | 71 | 263 | 1.1E-26 |
| TIGRFAM | TIGR01845 | outer_NodT: efflux transporter, outer membrane factor (OMF) lipoprotein, NodT family | IPR010131 | RND efflux system, outer membrane lipoprotein, NodT | 16 | 476 | 9.5E-120 |
| SUPERFAMILY | SSF56954 | 9 | 478 | 2.09E-102 | |||
| Coils | Coil | 230 | 264 | - | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 502 | 521 | - | ||
| Gene3D | G3DSA:2.20.200.10 | 108 | 365 | 7.7E-114 | |||
| ProSiteProfiles | PS51257 | Prokaryotic membrane lipoprotein lipid attachment site profile. | 1 | 24 | 5.0 | ||
| Gene3D | G3DSA:1.20.1600.10 | 61 | 476 | 7.7E-114 | |||
| Pfam | PF02321 | Outer membrane efflux protein | IPR003423 | Outer membrane efflux protein | 294 | 474 | 1.8E-23 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.