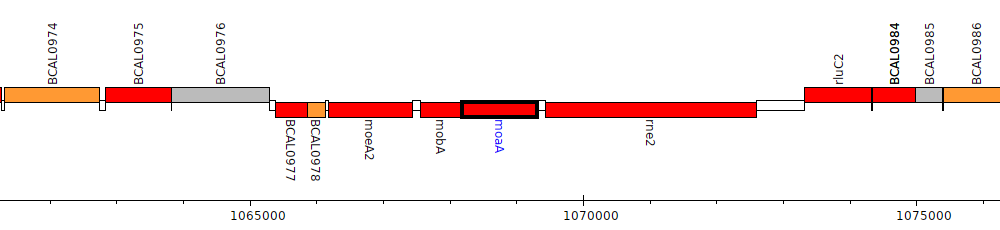
Burkholderia cenocepacia J2315, BCAL0981 (moaA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00729
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00729
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051539 | 4 iron, 4 sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS01305
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02666
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006777 | Mo-molybdopterin cofactor biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02666
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0019008 | molybdopterin synthase complex |
Inferred from Sequence Model
Term mapped from: InterPro:PF06463
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00790 | Folate biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj04122 | Sulfur relay system | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00790 | Folate biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | molybdenum cofactor biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Hamap | MF_01225_B | GTP 3',8-cyclase [moaA]. | IPR013483 | Molybdenum cofactor biosynthesis protein A | 28 | 370 | 40.34 |
| ProSitePatterns | PS01305 | moaA / nifB / pqqE family signature. | IPR000385 | MoaA/nifB/pqqE, iron-sulphur binding, conserved site | 43 | 54 | - |
| Pfam | PF06463 | Molybdenum Cofactor Synthesis C | IPR010505 | Molybdenum cofactor synthesis C-terminal | 218 | 344 | 1.7E-36 |
| SMART | SM00729 | IPR006638 | Elp3/MiaB/NifB | 37 | 251 | 3.2E-12 | |
| TIGRFAM | TIGR02666 | moaA: molybdenum cofactor biosynthesis protein A | IPR013483 | Molybdenum cofactor biosynthesis protein A | 28 | 370 | 9.6E-117 |
| Pfam | PF13353 | 4Fe-4S single cluster domain | 45 | 136 | 7.6E-8 | ||
| Gene3D | G3DSA:3.20.20.70 | IPR013785 | Aldolase-type TIM barrel | 25 | 370 | 9.7E-95 | |
| Pfam | PF04055 | Radical SAM superfamily | IPR007197 | Radical SAM | 41 | 210 | 9.8E-29 |
| CDD | cd01335 | Radical_SAM | 41 | 244 | 5.65114E-25 | ||
| SFLD | SFLDG01383 | cyclic pyranopterin phosphate synthase (MoaA-like) | 27 | 370 | 0.0 | ||
| SFLD | SFLDG01072 | dehydrogenase like | 27 | 370 | 0.0 | ||
| SUPERFAMILY | SSF102114 | 29 | 310 | 3.53E-73 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.