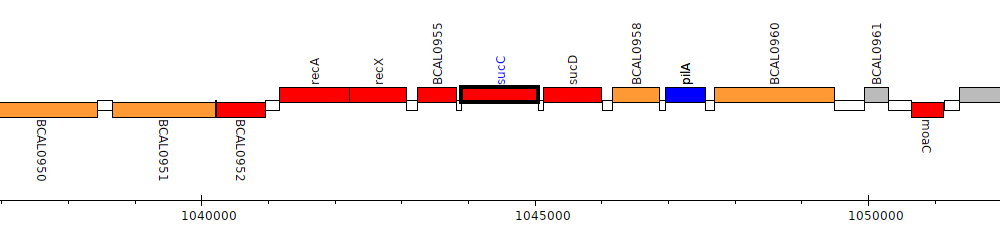
Burkholderia cenocepacia J2315, BCAL0956 (sucC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006099 | tricarboxylic acid cycle |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01016
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00549
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.30.1490.20
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS50975
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | TCA cycle II (plants and fungi) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | reductive TCA cycle II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | pyruvate fermentation to acetate V | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00640 | Propanoate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | partial TCA cycle (obligate autotrophs) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00720 | Carbon fixation pathways in prokaryotes | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | pyruvate fermentation to acetate VI | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | methylaspartate cycle | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00660 | C5-Branched dibasic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00020 | Citrate cycle (TCA cycle) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | TCA cycle V (2-oxoglutarate:ferredoxin oxidoreductase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00660 | C5-Branched dibasic acid metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | anaerobic energy metabolism (invertebrates, mitochondrial) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00640 | Propanoate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00020 | Citrate cycle (TCA cycle) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00549 | CoA-ligase | IPR005811 | ATP-citrate lyase/succinyl-CoA ligase | 262 | 382 | 1.5E-23 |
| Gene3D | G3DSA:3.30.1490.20 | IPR013815 | ATP-grasp fold, subdomain 1 | 21 | 104 | 2.0E-95 | |
| ProSitePatterns | PS01217 | ATP-citrate lyase / succinyl-CoA ligases family signature 3. | IPR017866 | Succinyl-CoA synthetase, beta subunit, conserved site | 257 | 282 | - |
| SUPERFAMILY | SSF52210 | IPR016102 | Succinyl-CoA synthetase-like | 239 | 384 | 1.44E-53 | |
| PIRSF | PIRSF001554 | IPR005809 | Succinate--CoA synthetase, beta subunit | 1 | 388 | 9.2E-154 | |
| Pfam | PF08442 | ATP-grasp domain | IPR013650 | ATP-grasp fold, succinyl-CoA synthetase-type | 2 | 202 | 3.8E-80 |
| Hamap | MF_00558 | Succinate--CoA ligase [ADP-forming] subunit beta [sucC]. | IPR005809 | Succinate--CoA synthetase, beta subunit | 1 | 386 | 43.634 |
| SUPERFAMILY | SSF56059 | 1 | 238 | 3.53E-83 | |||
| ProSiteProfiles | PS50975 | ATP-grasp fold profile. | IPR011761 | ATP-grasp fold | 9 | 231 | 15.652 |
| TIGRFAM | TIGR01016 | sucCoAbeta: succinate-CoA ligase, beta subunit | IPR005809 | Succinate--CoA synthetase, beta subunit | 1 | 385 | 2.1E-157 |
| Gene3D | G3DSA:3.30.470.20 | 1 | 234 | 2.0E-95 | |||
| Gene3D | G3DSA:3.40.50.261 | IPR016102 | Succinyl-CoA synthetase-like | 240 | 388 | 4.7E-57 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.