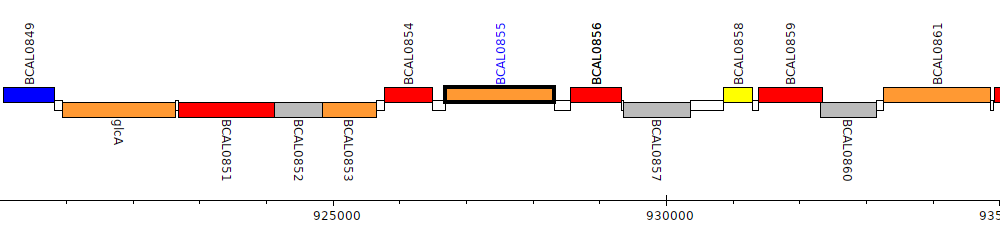
Burkholderia cenocepacia J2315, BCAL0855
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS50035
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.30.870.10 | 321 | 494 | 9.3E-36 | |||
| SMART | SM00155 | IPR001736 | Phospholipase D/Transphosphatidylase | 425 | 452 | 0.0042 | |
| SUPERFAMILY | SSF56024 | 331 | 481 | 1.14E-28 | |||
| CDD | cd09111 | PLDc_ymdC_like_1 | 81 | 246 | 1.8486E-76 | ||
| Gene3D | G3DSA:3.30.870.10 | 77 | 275 | 3.5E-36 | |||
| ProSiteProfiles | PS50035 | Phospholipase D phosphodiesterase active site profile. | IPR001736 | Phospholipase D/Transphosphatidylase | 186 | 213 | 13.112 |
| CDD | cd09113 | PLDc_ymdC_like_2 | 314 | 536 | 1.32659E-79 | ||
| Pfam | PF13091 | PLD-like domain | IPR025202 | Phospholipase D-like domain | 340 | 478 | 1.8E-23 |
| ProSiteProfiles | PS50035 | Phospholipase D phosphodiesterase active site profile. | IPR001736 | Phospholipase D/Transphosphatidylase | 425 | 452 | 12.638 |
| Pfam | PF13091 | PLD-like domain | IPR025202 | Phospholipase D-like domain | 95 | 245 | 2.3E-14 |
| SMART | SM00155 | IPR001736 | Phospholipase D/Transphosphatidylase | 186 | 213 | 3.8E-5 | |
| SUPERFAMILY | SSF56024 | 58 | 264 | 1.22E-39 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.