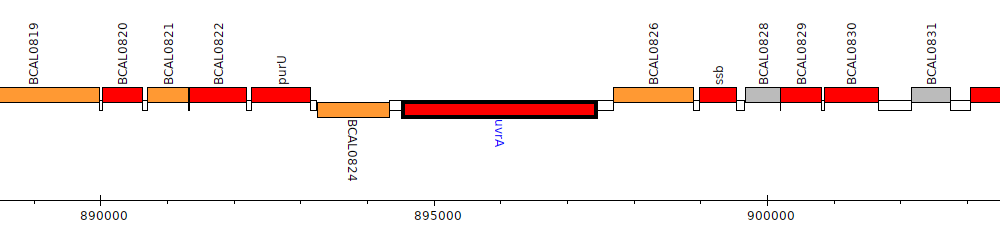
Burkholderia cenocepacia J2315, BCAL0825 (uvrA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00205
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00211
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00211
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009380 | excinuclease repair complex |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00205
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006289 | nucleotide-excision repair |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00205
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj03420 | Nucleotide excision repair | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.30.190.20 | 130 | 252 | 1.1E-243 | |||
| CDD | cd03271 | ABC_UvrA_II | 625 | 915 | 4.82645E-155 | ||
| CDD | cd03270 | ABC_UvrA_I | 4 | 116 | 1.10896E-74 | ||
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 631 | 924 | 5.06E-51 | |
| Pfam | PF17760 | UvrA interaction domain | IPR041102 | UvrA, interaction domain | 130 | 246 | 4.2E-31 |
| Gene3D | G3DSA:1.10.8.280 | 295 | 411 | 1.1E-243 | |||
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 1 | 581 | 1.16E-45 | |
| Pfam | PF17755 | UvrA DNA-binding domain | IPR041552 | UvrA DNA-binding domain | 298 | 407 | 4.1E-33 |
| TIGRFAM | TIGR00630 | uvra: excinuclease ABC subunit A | IPR004602 | UvrABC system subunit A | 3 | 936 | 0.0 |
| Gene3D | G3DSA:3.40.50.300 | 6 | 607 | 1.1E-243 | |||
| Gene3D | G3DSA:3.40.50.300 | 626 | 947 | 4.6E-141 | |||
| ProSitePatterns | PS00211 | ABC transporters family signature. | IPR017871 | ABC transporter, conserved site | 841 | 855 | - |
| ProSiteProfiles | PS50893 | ATP-binding cassette, ABC transporter-type domain profile. | IPR003439 | ABC transporter-like | 619 | 948 | 10.098 |
| CDD | cd03270 | ABC_UvrA_I | 358 | 587 | 1.12455E-70 | ||
| Hamap | MF_00205 | UvrABC system protein A [uvrA]. | IPR004602 | UvrABC system subunit A | 1 | 951 | 22.351 |
| Gene3D | G3DSA:1.20.1580.10 | 90 | 504 | 1.1E-243 | |||
| Gene3D | G3DSA:1.20.1580.10 | 704 | 841 | 4.6E-141 | |||
| ProSitePatterns | PS00211 | ABC transporters family signature. | IPR017871 | ABC transporter, conserved site | 499 | 513 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.