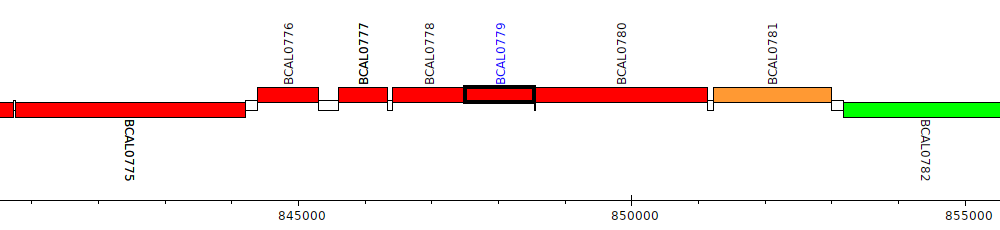
Burkholderia cenocepacia J2315, BCAL0779
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0097367 | carbohydrate derivative binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS51464
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:1901135 | carbohydrate derivative metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PS51464
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00250 | Alanine, aspartate and glutamate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | UDP-<i>N</i>-acetyl-D-galactosamine biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | CMP-legionaminate biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00520 | Amino sugar and nucleotide sugar metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00520 | Amino sugar and nucleotide sugar metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00250 | Alanine, aspartate and glutamate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF01380 | SIS domain | IPR001347 | Sugar isomerase (SIS) | 196 | 279 | 3.9E-14 |
| ProSiteProfiles | PS51464 | SIS domain profile. | IPR001347 | Sugar isomerase (SIS) | 192 | 336 | 18.242 |
| CDD | cd05009 | SIS_GlmS_GlmD_2 | IPR035490 | GlmS/FrlB, SIS domain 2 | 188 | 344 | 4.24778E-47 |
| Gene3D | G3DSA:3.40.50.10490 | 191 | 331 | 7.3E-39 | |||
| ProSiteProfiles | PS51464 | SIS domain profile. | IPR001347 | Sugar isomerase (SIS) | 29 | 176 | 18.646 |
| Pfam | PF01380 | SIS domain | IPR001347 | Sugar isomerase (SIS) | 42 | 160 | 4.3E-13 |
| CDD | cd05008 | SIS_GlmS_GlmD_1 | IPR035466 | GlmS/AgaS, SIS domain 1 | 41 | 164 | 1.16962E-32 |
| Gene3D | G3DSA:3.40.50.10490 | 1 | 187 | 2.4E-36 | |||
| SUPERFAMILY | SSF53697 | 3 | 344 | 4.58E-67 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.