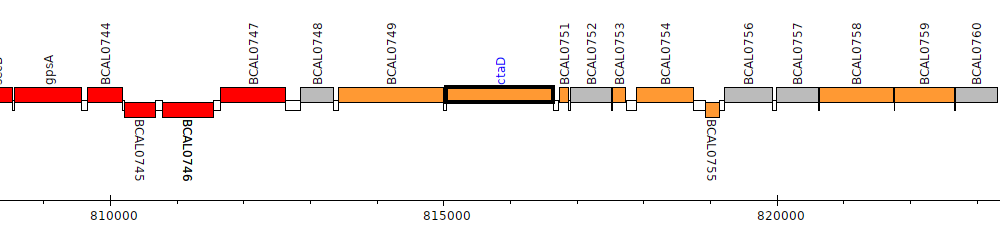
Burkholderia cenocepacia J2315, BCAL0750 (ctaD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009060 | aerobic respiration |
Inferred from Sequence Model
Term mapped from: InterPro:PR01165
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0020037 | heme binding |
Inferred from Sequence Model
Term mapped from: InterPro:PR01165
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0020037 | heme binding |
Inferred from Sequence Model
Term mapped from: InterPro:PR01165
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02891
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02891
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009060 | aerobic respiration |
Inferred from Sequence Model
Term mapped from: InterPro:PR01165
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0045277 | respiratory chain complex IV |
Inferred from Sequence Model
Term mapped from: InterPro:cd01663
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0045277 | respiratory chain complex IV |
Inferred from Sequence Model
Term mapped from: InterPro:cd01663
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004129 | cytochrome-c oxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02891
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0015990 | electron transport coupled proton transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02891
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0015990 | electron transport coupled proton transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02891
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PR01165
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004129 | cytochrome-c oxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02891
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PR01165
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00190 | Oxidative phosphorylation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 144 | 156 | 4.3E-120 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 74 | 97 | 4.3E-120 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 436 | 457 | 4.3E-120 |
| Pfam | PF00115 | Cytochrome C and Quinol oxidase polypeptide I | IPR000883 | Cytochrome c oxidase subunit I | 36 | 477 | 1.6E-152 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 101 | 125 | 4.3E-120 |
| ProSiteProfiles | PS50855 | Cytochrome oxidase subunit I profile. | IPR023616 | Cytochrome c oxidase-like, subunit I domain | 19 | 535 | 104.52 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 27 | 52 | 4.3E-120 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 386 | 405 | 4.3E-120 |
| SUPERFAMILY | SSF81442 | IPR036927 | Cytochrome c oxidase-like, subunit I superfamily | 24 | 534 | 9.29E-190 | |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 299 | 314 | 4.3E-120 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 253 | 274 | 4.3E-120 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 358 | 376 | 4.3E-120 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 173 | 191 | 4.3E-120 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 323 | 344 | 4.3E-120 |
| Gene3D | G3DSA:1.20.210.10 | IPR036927 | Cytochrome c oxidase-like, subunit I superfamily | 15 | 535 | 2.6E-216 | |
| TIGRFAM | TIGR02891 | CtaD_CoxA: cytochrome c oxidase, subunit I | IPR014241 | Cytochrome c oxidase, subunit I bacterial type | 28 | 534 | 1.5E-211 |
| ProSitePatterns | PS00077 | Heme-copper oxidase catalytic subunit, copper B binding region signature. | IPR023615 | Cytochrome c oxidase, subunit I, copper-binding site | 255 | 309 | - |
| CDD | cd01663 | Cyt_c_Oxidase_I | IPR033944 | Cytochrome c oxidase subunit I domain | 32 | 519 | 0.0 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 202 | 221 | 4.3E-120 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.