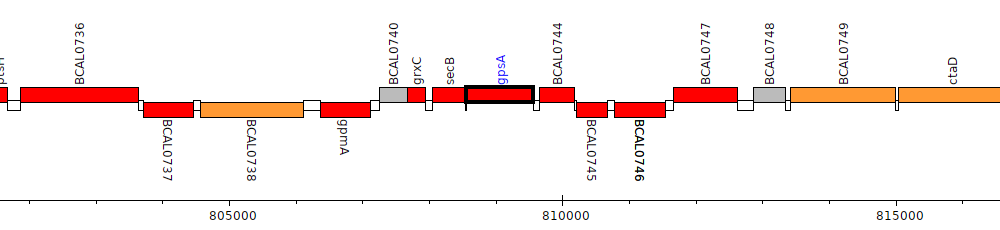
Burkholderia cenocepacia J2315, BCAL0743 (gpsA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004367 | glycerol-3-phosphate dehydrogenase [NAD+] activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07479
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0046168 | glycerol-3-phosphate catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF01210
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PF01210
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009331 | glycerol-3-phosphate dehydrogenase complex |
Inferred from Sequence Model
Term mapped from: InterPro:PR00077
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.10.1040.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF07479
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006072 | glycerol-3-phosphate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PR00077
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051287 | NAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01210
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF07479
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00564 | Glycerophospholipid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | CDP-diacylglycerol biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | glucosylglycerol biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00564 | Glycerophospholipid metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | CDP-diacylglycerol biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00077 | NAD-dependent glycerol-3-phosphate dehydrogenase signature | IPR006168 | Glycerol-3-phosphate dehydrogenase, NAD-dependent | 239 | 256 | 1.1E-35 |
| PRINTS | PR00077 | NAD-dependent glycerol-3-phosphate dehydrogenase signature | IPR006168 | Glycerol-3-phosphate dehydrogenase, NAD-dependent | 134 | 154 | 1.1E-35 |
| Coils | Coil | 326 | 332 | - | |||
| SUPERFAMILY | SSF51735 | IPR036291 | NAD(P)-binding domain superfamily | 1 | 181 | 2.92E-35 | |
| PRINTS | PR00077 | NAD-dependent glycerol-3-phosphate dehydrogenase signature | IPR006168 | Glycerol-3-phosphate dehydrogenase, NAD-dependent | 199 | 223 | 1.1E-35 |
| Pfam | PF07479 | NAD-dependent glycerol-3-phosphate dehydrogenase C-terminus | IPR006109 | Glycerol-3-phosphate dehydrogenase, NAD-dependent, C-terminal | 181 | 321 | 1.2E-47 |
| PIRSF | PIRSF000114 | IPR006168 | Glycerol-3-phosphate dehydrogenase, NAD-dependent | 1 | 332 | 2.4E-92 | |
| ProSitePatterns | PS00957 | NAD-dependent glycerol-3-phosphate dehydrogenase signature. | IPR006168 | Glycerol-3-phosphate dehydrogenase, NAD-dependent | 189 | 210 | - |
| SUPERFAMILY | SSF48179 | IPR008927 | 6-phosphogluconate dehydrogenase-like, C-terminal domain superfamily | 180 | 332 | 1.51E-51 | |
| Pfam | PF01210 | NAD-dependent glycerol-3-phosphate dehydrogenase N-terminus | IPR011128 | Glycerol-3-phosphate dehydrogenase, NAD-dependent, N-terminal | 2 | 158 | 5.7E-37 |
| Hamap | MF_00394 | Glycerol-3-phosphate dehydrogenase [NAD(P)+] [gpsA]. | IPR006168 | Glycerol-3-phosphate dehydrogenase, NAD-dependent | 1 | 323 | 34.336 |
| Gene3D | G3DSA:3.40.50.720 | 1 | 188 | 2.1E-48 | |||
| PRINTS | PR00077 | NAD-dependent glycerol-3-phosphate dehydrogenase signature | IPR006168 | Glycerol-3-phosphate dehydrogenase, NAD-dependent | 174 | 198 | 1.1E-35 |
| PRINTS | PR00077 | NAD-dependent glycerol-3-phosphate dehydrogenase signature | IPR006168 | Glycerol-3-phosphate dehydrogenase, NAD-dependent | 5 | 22 | 1.1E-35 |
| Gene3D | G3DSA:1.10.1040.10 | IPR013328 | 6-phosphogluconate dehydrogenase, domain 2 | 190 | 332 | 7.4E-46 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.