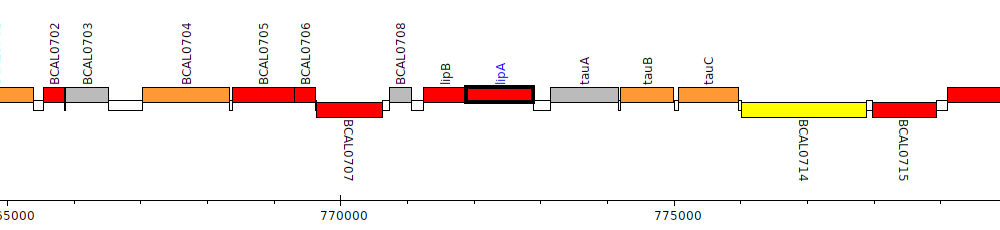
Burkholderia cenocepacia J2315, BCAL0710 (lipA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDS00029
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016992 | lipoate synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDF00271
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDS00029
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051539 | 4 iron, 4 sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDF00271
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009107 | lipoate biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDF00271
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00785 | Lipoic acid metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | lipoate biosynthesis and incorporation III (Bacillus) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00785 | Lipoic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | lipoate biosynthesis and incorporation (yeast) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 23 | - | ||
| SUPERFAMILY | SSF102114 | 60 | 325 | 2.88E-68 | |||
| SMART | SM00729 | IPR006638 | Elp3/MiaB/NifB | 93 | 302 | 6.3E-19 | |
| SFLD | SFLDF00271 | lipoyl synthase | IPR003698 | Lipoyl synthase | 42 | 324 | 0.0 |
| PIRSF | PIRSF005963 | IPR003698 | Lipoyl synthase | 5 | 328 | 2.9E-133 | |
| SFLD | SFLDS00029 | Radical SAM | IPR007197 | Radical SAM | 42 | 324 | 0.0 |
| CDD | cd01335 | Radical_SAM | 103 | 295 | 7.0383E-11 | ||
| Pfam | PF04055 | Radical SAM superfamily | IPR007197 | Radical SAM | 100 | 260 | 7.2E-17 |
| Gene3D | G3DSA:3.20.20.70 | IPR013785 | Aldolase-type TIM barrel | 89 | 303 | 3.3E-21 | |
| Hamap | MF_00206 | Lipoyl synthase [lipA]. | IPR003698 | Lipoyl synthase | 45 | 330 | 155.306 |
| Pfam | PF16881 | N-terminal domain of lipoyl synthase of Radical_SAM family | IPR031691 | Lipoyl synthase, N-terminal | 41 | 82 | 3.6E-9 |
| TIGRFAM | TIGR00510 | lipA: lipoyl synthase | IPR003698 | Lipoyl synthase | 42 | 325 | 4.9E-132 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.