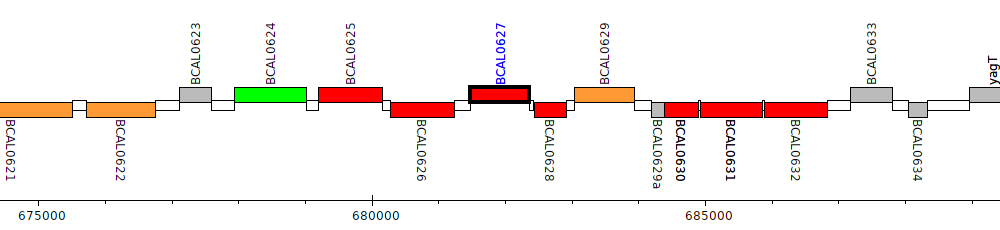
Burkholderia cenocepacia J2315, BCAL0627
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00361 | Chlorocyclohexane and chlorobenzene degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00625 | Chloroalkane and chloroalkene degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.1820 | IPR029058 | Alpha/Beta hydrolase fold | 2 | 293 | 1.2E-83 | |
| Pfam | PF00561 | alpha/beta hydrolase fold | IPR000073 | Alpha/beta hydrolase fold-1 | 27 | 152 | 5.9E-21 |
| SUPERFAMILY | SSF53474 | IPR029058 | Alpha/Beta hydrolase fold | 5 | 292 | 5.14E-71 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.