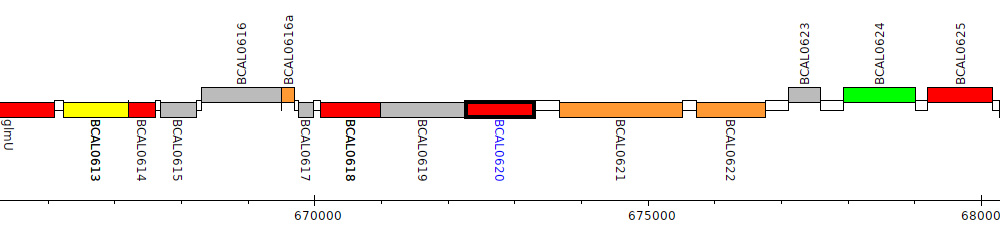
Burkholderia cenocepacia J2315, BCAL0620
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF47413
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:cd01392
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00354 | IPR000843 | LacI-type HTH domain | 2 | 57 | 2.0E-6 | |
| Gene3D | G3DSA:1.10.260.40 | 1 | 57 | 6.9E-18 | |||
| CDD | cd06267 | PBP1_LacI_sugar_binding_like | 71 | 318 | 2.93148E-67 | ||
| Pfam | PF13377 | Periplasmic binding protein-like domain | 163 | 323 | 2.4E-32 | ||
| SUPERFAMILY | SSF47413 | IPR010982 | Lambda repressor-like, DNA-binding domain superfamily | 3 | 51 | 5.91E-13 | |
| Pfam | PF00356 | Bacterial regulatory proteins, lacI family | IPR000843 | LacI-type HTH domain | 4 | 48 | 1.3E-14 |
| SUPERFAMILY | SSF53822 | IPR028082 | Periplasmic binding protein-like I | 71 | 308 | 1.81E-54 | |
| Gene3D | G3DSA:3.40.50.2300 | 72 | 315 | 1.2E-59 | |||
| Gene3D | G3DSA:3.40.50.2300 | 154 | 320 | 1.2E-59 | |||
| CDD | cd01392 | HTH_LacI | IPR000843 | LacI-type HTH domain | 6 | 57 | 6.10526E-15 |
| ProSitePatterns | PS00356 | LacI-type HTH domain signature. | 5 | 23 | - | ||
| ProSiteProfiles | PS50932 | LacI-type HTH domain profile. | IPR000843 | LacI-type HTH domain | 3 | 57 | 25.012 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.