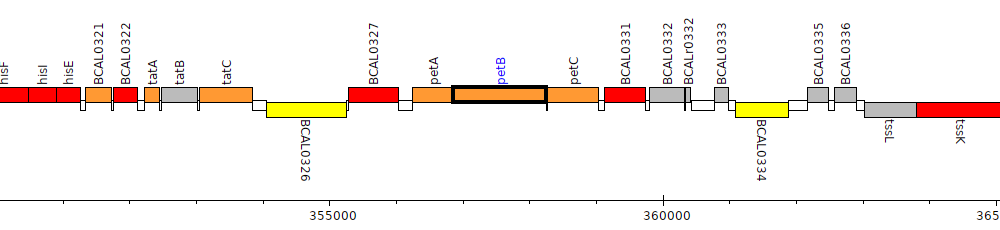
Burkholderia cenocepacia J2315, BCAL0329 (petB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00032
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF00032
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008121 | ubiquinol-cytochrome-c reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF038885
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006122 | mitochondrial electron transport, ubiquinol to cytochrome c |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF038885
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0022904 | respiratory electron transport chain |
Inferred from Sequence Model
Term mapped from: InterPro:SSF81342
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0045275 | respiratory chain complex III |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF038885
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00032
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj02020 | Two-component system | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00190 | Oxidative phosphorylation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00032 | Cytochrome b(C-terminal)/b6/petD | IPR005798 | Cytochrome b/b6, C-terminal | 286 | 433 | 3.0E-38 |
| ProSiteProfiles | PS51002 | Cytochrome b/b6 N-terminal region profile. | IPR005797 | Cytochrome b/b6, N-terminal | 13 | 225 | 39.559 |
| PIRSF | PIRSF038885 | IPR030689 | Cytochrome b | 5 | 457 | 3.7E-152 | |
| Pfam | PF00033 | Cytochrome b/b6/petB | IPR005797 | Cytochrome b/b6, N-terminal | 32 | 220 | 1.7E-66 |
| CDD | cd00284 | Cytochrome_b_N | IPR005797 | Cytochrome b/b6, N-terminal | 22 | 222 | 4.16582E-72 |
| Gene3D | G3DSA:1.20.810.10 | IPR027387 | Cytochrome b/b6-like domain superfamily | 354 | 448 | 1.1E-15 | |
| ProSiteProfiles | PS51003 | Cytochrome b/b6 C-terminal region profile. | IPR005798 | Cytochrome b/b6, C-terminal | 235 | 454 | 33.057 |
| Gene3D | G3DSA:1.20.810.10 | IPR027387 | Cytochrome b/b6-like domain superfamily | 2 | 353 | 4.1E-130 | |
| SUPERFAMILY | SSF81648 | IPR036150 | Cytochrome b/b6, C-terminal domain superfamily | 289 | 443 | 1.83E-21 | |
| SUPERFAMILY | SSF81342 | IPR016174 | Di-haem cytochrome, transmembrane | 16 | 288 | 1.88E-97 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.