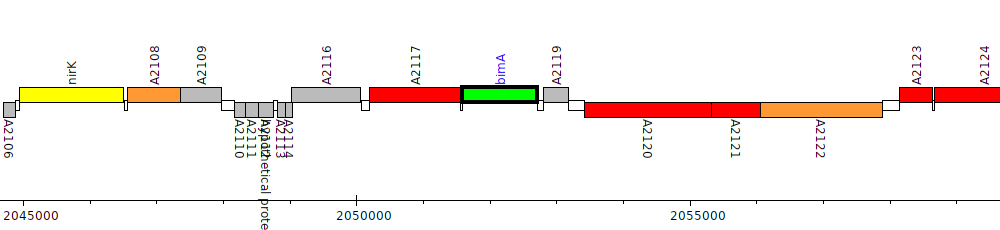
Burkholderia pseudomallei 668, BURPS668_A2118 (bimA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009405 | pathogenesis |
Inferred from Sequence Model
Term mapped from: InterPro:PF05662
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0019867 | outer membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF05662
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| MobiDBLite | mobidb-lite | consensus disorder prediction | 85 | 138 | - | ||
| Pfam | PF03895 | YadA-like membrane anchor domain | IPR005594 | YadA-like, C-terminal | 320 | 378 | 3.6E-16 |
| Gene3D | G3DSA:3.30.1300.30 | 281 | 378 | 6.8E-21 | |||
| SUPERFAMILY | SSF54523 | 269 | 378 | 6.59E-22 | |||
| Pfam | PF05662 | Coiled stalk of trimeric autotransporter adhesin | IPR008635 | Coiled stalk of trimeric autotransporter adhesin | 258 | 295 | 5.3E-10 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 83 | 152 | - | ||
| SUPERFAMILY | SSF101967 | 179 | 278 | 6.41E-23 | |||
| PRINTS | PR01217 | Proline rich extensin signature | 90 | 106 | 1.3E-11 | ||
| PRINTS | PR01217 | Proline rich extensin signature | 110 | 122 | 1.3E-11 | ||
| PRINTS | PR01217 | Proline rich extensin signature | 125 | 146 | 1.3E-11 | ||
| Gene3D | G3DSA:2.150.10.10 | IPR011049 | Serralysin-like metalloprotease, C-terminal | 163 | 280 | 6.4E-27 | |
| Pfam | PF05658 | Head domain of trimeric autotransporter adhesin | IPR008640 | Head domain of trimeric autotransporter adhesin | 228 | 251 | 5.4E-4 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.