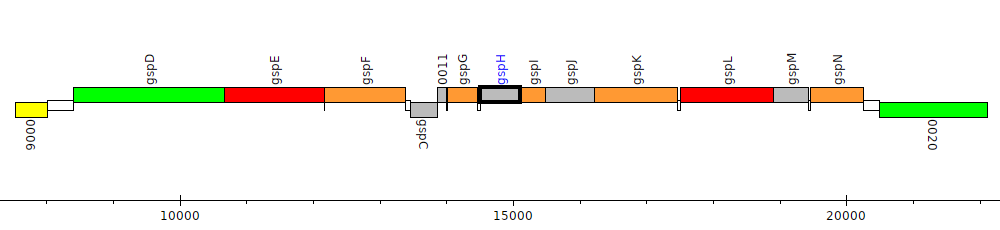
Burkholderia pseudomallei 1106a, BURPS1106A_0013 (gspH)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0015628 | protein secretion by the type II secretion system | Inferred from Genomic Context | ECO:0000177 genomic context evidence |
||
| Biological Process | GO:0015628 | protein secretion by the type II secretion system |
Inferred from Sequence Model
Term mapped from: InterPro:PF12019
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0015627 | type II protein secretion system complex |
Inferred from Sequence Model
Term mapped from: InterPro:PF12019
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bpl03070 | Bacterial secretion system | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF12019 | Type II transport protein GspH | IPR022346 | General secretion pathway, GspH | 89 | 194 | 3.4E-7 |
| TIGRFAM | TIGR01708 | typeII_sec_gspH: type II secretion system protein H | IPR002416 | Type II secretion system protein H | 50 | 197 | 4.8E-27 |
| SUPERFAMILY | SSF54523 | 55 | 136 | 9.42E-12 | |||
| Pfam | PF07963 | Prokaryotic N-terminal methylation motif | IPR012902 | Prokaryotic N-terminal methylation site | 50 | 74 | 1.3E-8 |
| Gene3D | G3DSA:3.55.40.10 | 83 | 169 | 2.1E-9 | |||
| ProSitePatterns | PS00409 | Prokaryotic N-terminal methylation site. | IPR012902 | Prokaryotic N-terminal methylation site | 53 | 73 | - |
| TIGRFAM | TIGR02532 | IV_pilin_GFxxxE: prepilin-type N-terminal cleavage/methylation domain | IPR012902 | Prokaryotic N-terminal methylation site | 53 | 74 | 2.1E-8 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.