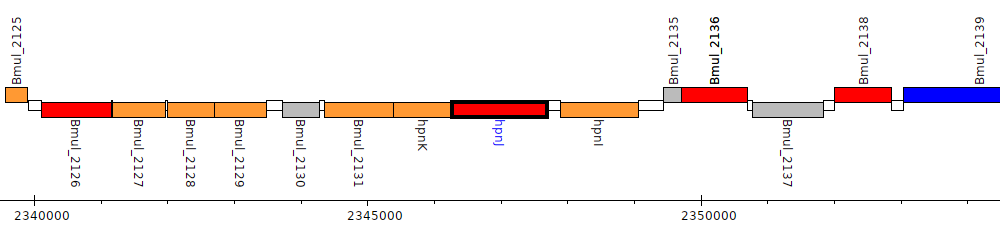
Burkholderia multivorans ATCC 17616, Bmul_2133 (hpnJ)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0090559 | regulation of membrane permeability | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
22006009 | Reviewed by curator |
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDS00029
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDS00029
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SFLD | SFLDG01123 | methyltransferase (Class B) | IPR034466 | Methyltransferase, class B | 1 | 439 | 0.0 |
| Pfam | PF04055 | Radical SAM superfamily | IPR007197 | Radical SAM | 202 | 361 | 4.2E-19 |
| SFLD | SFLDF00404 | hopanetetrol cyclitol ether synthase | IPR017834 | Hopanoid biosynthesis associated radical SAM protein HpnJ | 1 | 472 | 0.0 |
| SUPERFAMILY | SSF102114 | 190 | 467 | 1.31E-44 | |||
| TIGRFAM | TIGR03471 | HpnJ: hopanoid biosynthesis associated radical SAM protein HpnJ | IPR017834 | Hopanoid biosynthesis associated radical SAM protein HpnJ | 1 | 472 | 7.2E-253 |
| SMART | SM00729 | IPR006638 | Elp3/MiaB/NifB | 196 | 404 | 3.8E-27 | |
| Gene3D | G3DSA:3.80.30.20 | IPR023404 | Radical SAM, alpha/beta horseshoe | 194 | 405 | 2.0E-26 | |
| SFLD | SFLDS00029 | Radical SAM | IPR007197 | Radical SAM | 1 | 472 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.