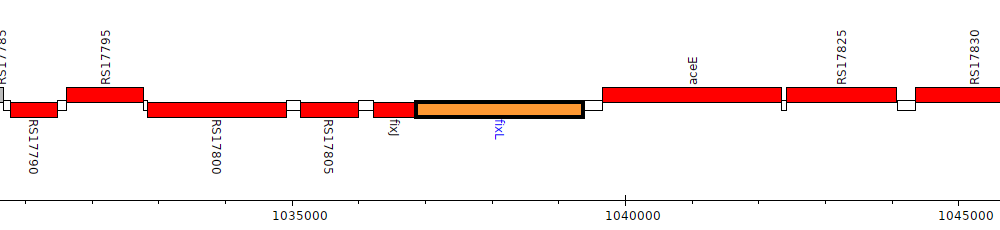
Burkholderia dolosa AU0158, AK34_RS17815 (fixL)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0036294 | cellular response to decreased oxygen levels |
Inferred from Genetic Interaction
Term mapped from: locus_tag:AK34_RS17810
|
ECO:0000054 double mutant phenotype evidence |
28046077 | Reviewed by curator |
| Biological Process | GO:0000160 | phosphorelay signal transduction system |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:P10955
|
ECO:0000250 sequence similarity evidence used in manual assertion |
28046077 | Reviewed by curator |
| Molecular Function | GO:0000155 | phosphorelay sensor kinase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:P10955
|
ECO:0000250 sequence similarity evidence used in manual assertion |
28046077 | Reviewed by curator |
| Biological Process | GO:0016310 | phosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:PR00344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PF00989
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000155 | phosphorelay sensor kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00512
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:PF00512
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016772 | transferase activity, transferring phosphorus-containing groups |
Inferred from Sequence Model
Term mapped from: InterPro:PR00344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF55785 | IPR035965 | PAS domain superfamily | 433 | 564 | 4.36E-9 | |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 743 | 757 | 5.1E-13 |
| Gene3D | G3DSA:3.30.450.20 | 446 | 580 | 1.8E-7 | |||
| TIGRFAM | TIGR00229 | sensory_box: PAS domain S-box protein | IPR000014 | PAS domain | 318 | 443 | 1.6E-24 |
| ProSiteProfiles | PS50109 | Histidine kinase domain profile. | IPR005467 | Histidine kinase domain | 595 | 817 | 37.865 |
| ProSiteProfiles | PS50113 | PAC domain profile. | IPR000700 | PAS-associated, C-terminal | 393 | 445 | 14.681 |
| CDD | cd00130 | PAS | IPR000014 | PAS domain | 332 | 433 | 9.54817E-15 |
| Pfam | PF00512 | His Kinase A (phospho-acceptor) domain | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 589 | 655 | 3.0E-7 |
| Pfam | PF00989 | PAS fold | IPR013767 | PAS fold | 322 | 433 | 1.5E-15 |
| SMART | SM00091 | IPR000014 | PAS domain | 319 | 387 | 3.0E-7 | |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 772 | 790 | 5.1E-13 |
| SUPERFAMILY | SSF47384 | IPR036097 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain superfamily | 582 | 656 | 3.34E-8 | |
| SUPERFAMILY | SSF55874 | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 650 | 815 | 1.96E-34 | |
| SMART | SM00388 | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 588 | 658 | 2.9E-8 | |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 801 | 814 | 5.1E-13 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 761 | 771 | 5.1E-13 |
| Pfam | PF13188 | PAS domain | IPR000014 | PAS domain | 448 | 492 | 0.013 |
| Gene3D | G3DSA:3.30.565.10 | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 657 | 819 | 9.6E-36 | |
| SMART | SM00091 | IPR000014 | PAS domain | 448 | 518 | 64.0 | |
| SMART | SM00387 | IPR003594 | Histidine kinase/HSP90-like ATPase | 700 | 817 | 3.6E-28 | |
| CDD | cd16920 | HATPase_TmoS-FixL-DctS-like | 705 | 814 | 2.72777E-39 | ||
| SMART | SM00086 | IPR001610 | PAC motif | 394 | 436 | 1.8E-6 | |
| ProSiteProfiles | PS50112 | PAS repeat profile. | IPR000014 | PAS domain | 317 | 389 | 15.35 |
| Gene3D | G3DSA:1.10.287.130 | 593 | 655 | 7.3E-18 | |||
| Pfam | PF02518 | Histidine kinase-, DNA gyrase B-, and HSP90-like ATPase | IPR003594 | Histidine kinase/HSP90-like ATPase | 702 | 814 | 9.5E-15 |
| CDD | cd00082 | HisKA | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 586 | 654 | 4.1717E-7 |
| Gene3D | G3DSA:3.30.450.20 | 317 | 445 | 1.1E-25 | |||
| SUPERFAMILY | SSF55785 | IPR035965 | PAS domain superfamily | 307 | 434 | 1.61E-24 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.