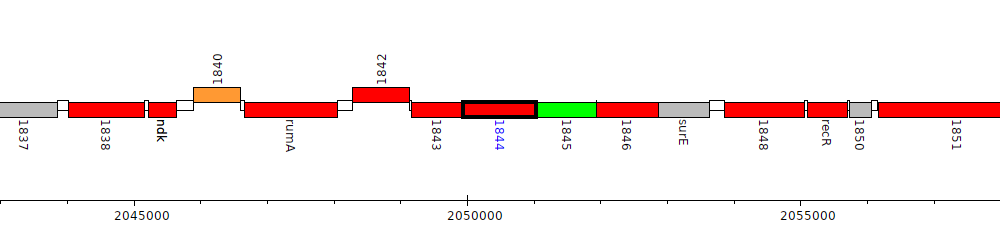
Burkholderia cenocepacia MC0-3, Bcenmc03_1844
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016987 | sigma factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016987 | sigma factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006352 | DNA-templated transcription, initiation |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006352 | DNA-templated transcription, initiation |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003700 | DNA-binding transcription factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003700 | DNA-binding transcription factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00959
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR02937 | sigma70-ECF: RNA polymerase sigma factor, sigma-70 family | IPR014284 | RNA polymerase sigma-70 like domain | 115 | 349 | 1.0E-33 |
| Pfam | PF04542 | Sigma-70 region 2 | IPR007627 | RNA polymerase sigma-70 region 2 | 120 | 189 | 8.5E-20 |
| SUPERFAMILY | SSF88659 | IPR013324 | RNA polymerase sigma factor, region 3/4-like | 193 | 275 | 3.88E-7 | |
| Gene3D | G3DSA:1.10.601.10 | 4 | 192 | 6.9E-50 | |||
| CDD | cd06171 | Sigma70_r4 | 291 | 348 | 2.86475E-9 | ||
| Hamap | MF_00959 | RNA polymerase sigma factor RpoS [rpoS]. | IPR012761 | RNA polymerase sigma factor RpoS | 3 | 362 | 38.33 |
| TIGRFAM | TIGR02394 | rpoS_proteo: RNA polymerase sigma factor RpoS | IPR012761 | RNA polymerase sigma factor RpoS | 73 | 361 | 4.7E-140 |
| ProSitePatterns | PS00715 | Sigma-70 factors family signature 1. | IPR000943 | RNA polymerase sigma-70 | 144 | 157 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 42 | 68 | - | ||
| Pfam | PF04545 | Sigma-70, region 4 | IPR007630 | RNA polymerase sigma-70 region 4 | 296 | 348 | 7.8E-17 |
| SUPERFAMILY | SSF88659 | IPR013324 | RNA polymerase sigma factor, region 3/4-like | 259 | 356 | 2.29E-24 | |
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 144 | 157 | 1.1E-23 |
| Gene3D | G3DSA:1.10.10.10 | IPR036388 | Winged helix-like DNA-binding domain superfamily | 278 | 353 | 1.5E-19 | |
| Gene3D | G3DSA:1.10.10.10 | IPR036388 | Winged helix-like DNA-binding domain superfamily | 193 | 276 | 5.0E-13 | |
| Pfam | PF04539 | Sigma-70 region 3 | IPR007624 | RNA polymerase sigma-70 region 3 | 201 | 282 | 5.0E-7 |
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 336 | 347 | 1.1E-23 |
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 300 | 312 | 1.1E-23 |
| Coils | Coil | 322 | 342 | - | |||
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 321 | 336 | 1.1E-23 |
| ProSitePatterns | PS00716 | Sigma-70 factors family signature 2. | IPR000943 | RNA polymerase sigma-70 | 321 | 347 | - |
| SUPERFAMILY | SSF88946 | IPR013325 | RNA polymerase sigma factor, region 2 | 82 | 190 | 4.08E-44 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 69 | - | ||
| Pfam | PF00140 | Sigma-70 factor, region 1.2 | IPR009042 | RNA polymerase sigma-70 region 1.2 | 82 | 115 | 4.2E-8 |
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 168 | 176 | 1.1E-23 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.