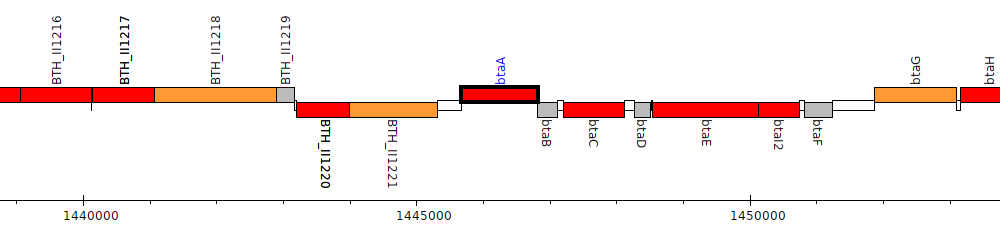
Burkholderia thailandensis E264 ATCC 700388, BTH_II1222 (btaA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0017000 | antibiotic biosynthetic process | Inferred from Genomic Context | ECO:0000317 genomic context evidence used in manual assertion |
20095633 | Reviewed by curator |
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF009283
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003868 | 4-hydroxyphenylpyruvate dioxygenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF009283
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016701 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF009283
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009072 | aromatic amino acid family metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF009283
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bte00350 | Tyrosine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00130 | Ubiquinone and other terpenoid-quinone biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00360 | Phenylalanine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF14696 | Hydroxyphenylpyruvate dioxygenase, HPPD, N-terminal | 20 | 166 | 8.0E-49 | ||
| ProSiteProfiles | PS51819 | Vicinal oxygen chelate (VOC) domain profile. | IPR037523 | Vicinal oxygen chelate (VOC) domain | 187 | 354 | 20.78 |
| PIRSF | PIRSF009283 | IPR005956 | 4-hydroxyphenylpyruvate dioxygenase | 12 | 381 | 1.5E-101 | |
| CDD | cd07250 | HPPD_C_like | IPR041735 | 4-hydroxyphenylpyruvate dioxygenase, C-terminal | 185 | 371 | 2.61003E-63 |
| ProSiteProfiles | PS51819 | Vicinal oxygen chelate (VOC) domain profile. | IPR037523 | Vicinal oxygen chelate (VOC) domain | 27 | 145 | 16.478 |
| Gene3D | G3DSA:3.10.180.10 | IPR029068 | Glyoxalase/Bleomycin resistance protein/Dihydroxybiphenyl dioxygenase | 183 | 379 | 2.5E-46 | |
| Gene3D | G3DSA:3.10.180.10 | IPR029068 | Glyoxalase/Bleomycin resistance protein/Dihydroxybiphenyl dioxygenase | 19 | 176 | 9.5E-44 | |
| SUPERFAMILY | SSF54593 | IPR029068 | Glyoxalase/Bleomycin resistance protein/Dihydroxybiphenyl dioxygenase | 20 | 368 | 9.19E-68 | |
| Pfam | PF00903 | Glyoxalase/Bleomycin resistance protein/Dioxygenase superfamily | IPR004360 | Glyoxalase/fosfomycin resistance/dioxygenase domain | 188 | 299 | 4.4E-15 |
| CDD | cd08342 | HPPD_N_like | IPR041736 | 4-hydroxyphenylpyruvate dioxygenase, N-terminal | 29 | 168 | 5.35369E-28 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.