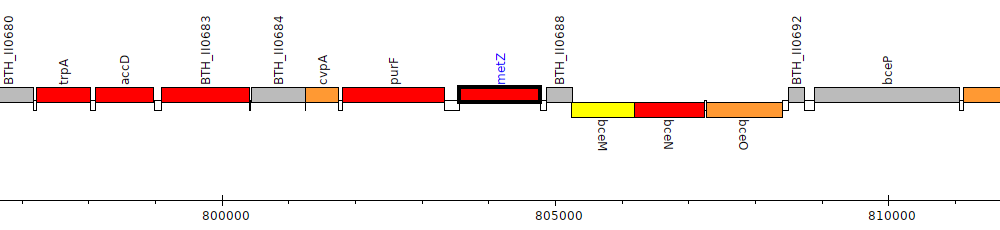
Burkholderia thailandensis E264 ATCC 700388, BTH_II0687 (metZ)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.640.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0071268 | homocysteine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_02056
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019346 | transsulfuration |
Inferred from Sequence Model
Term mapped from: InterPro:cd00614
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030170 | pyridoxal phosphate binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00614
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030170 | pyridoxal phosphate binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00614
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019346 | transsulfuration |
Inferred from Sequence Model
Term mapped from: InterPro:cd00614
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.640.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0071268 | homocysteine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_02056
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bte01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00270 | Cysteine and methionine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00920 | Sulfur metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF53383 | IPR015424 | Pyridoxal phosphate-dependent transferase | 38 | 404 | 1.75E-119 | |
| TIGRFAM | TIGR01325 | O_suc_HS_sulf: O-succinylhomoserine sulfhydrylase | IPR006234 | O-succinylhomoserine sulfhydrylase | 17 | 403 | 5.7E-174 |
| ProSitePatterns | PS00868 | Cys/Met metabolism enzymes pyridoxal-phosphate attachment site. | IPR000277 | Cys/Met metabolism, pyridoxal phosphate-dependent enzyme | 208 | 222 | - |
| Hamap | MF_02056 | O-succinylhomoserine sulfhydrylase [metZ]. | IPR006234 | O-succinylhomoserine sulfhydrylase | 18 | 404 | 55.053 |
| PIRSF | PIRSF001434 | IPR000277 | Cys/Met metabolism, pyridoxal phosphate-dependent enzyme | 3 | 405 | 9.6E-132 | |
| Gene3D | G3DSA:3.90.1150.10 | IPR015422 | Pyridoxal phosphate-dependent transferase domain 1 | 267 | 405 | 4.2E-50 | |
| Pfam | PF01053 | Cys/Met metabolism PLP-dependent enzyme | IPR000277 | Cys/Met metabolism, pyridoxal phosphate-dependent enzyme | 17 | 403 | 1.4E-137 |
| Gene3D | G3DSA:3.40.640.10 | IPR015421 | Pyridoxal phosphate-dependent transferase, major domain | 15 | 266 | 1.1E-85 | |
| CDD | cd00614 | CGS_like | IPR000277 | Cys/Met metabolism, pyridoxal phosphate-dependent enzyme | 37 | 404 | 5.09236E-160 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.