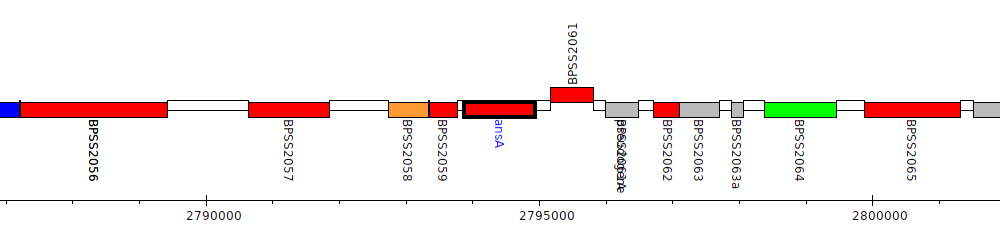
Burkholderia pseudomallei K96243, BPSS2060 (ansA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006520 | cellular amino acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PS00144
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004067 | asparaginase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00520
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006528 | asparagine metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00520
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00460 | Cyanoamino acid metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps00250 | Alanine, aspartate and glutamate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00250 | Alanine, aspartate and glutamate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00460 | Cyanoamino acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF53774 | IPR036152 | Asparaginase/glutaminase-like superfamily | 25 | 348 | 3.93E-98 | |
| Gene3D | G3DSA:3.40.50.1170 | IPR037152 | L-asparaginase, N-terminal domain superfamily | 23 | 231 | 6.0E-77 | |
| ProSiteProfiles | PS51732 | Asparaginase / glutaminase domain profile. | IPR006034 | Asparaginase/glutaminase-like | 26 | 350 | 84.426 |
| CDD | cd08964 | L-asparaginase_II | IPR004550 | L-asparaginase, type II | 26 | 344 | 1.64423E-137 |
| Pfam | PF17763 | Glutaminase/Asparaginase C-terminal domain | IPR040919 | Asparaginase/glutaminase, C-terminal | 236 | 347 | 7.3E-27 |
| PRINTS | PR00139 | Asparaginase/glutaminase family signature | IPR006034 | Asparaginase/glutaminase-like | 283 | 301 | 3.1E-23 |
| PIRSF | PIRSF001220 | IPR006034 | Asparaginase/glutaminase-like | 22 | 351 | 1.7E-93 | |
| SMART | SM00870 | IPR006034 | Asparaginase/glutaminase-like | 27 | 343 | 1.2E-129 | |
| PRINTS | PR00139 | Asparaginase/glutaminase family signature | IPR006034 | Asparaginase/glutaminase-like | 28 | 39 | 3.1E-23 |
| ProSitePatterns | PS00144 | Asparaginase / glutaminase active site signature 1. | IPR020827 | Asparaginase/glutaminase, active site 1 | 30 | 38 | - |
| TIGRFAM | TIGR00520 | asnASE_II: L-asparaginase, type II | IPR004550 | L-asparaginase, type II | 12 | 348 | 1.1E-116 |
| PRINTS | PR00139 | Asparaginase/glutaminase family signature | IPR006034 | Asparaginase/glutaminase-like | 107 | 125 | 3.1E-23 |
| Gene3D | G3DSA:3.40.50.40 | IPR027473 | L-asparaginase, C-terminal | 236 | 349 | 5.4E-37 | |
| PIRSF | PIRSF500176 | 24 | 351 | 1.8E-82 | |||
| ProSitePatterns | PS00917 | Asparaginase / glutaminase active site signature 2. | IPR027475 | Asparaginase/glutaminase, active site 2 | 108 | 118 | - |
| Pfam | PF00710 | Asparaginase, N-terminal | IPR027474 | L-asparaginase, N-terminal | 27 | 217 | 2.9E-59 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.