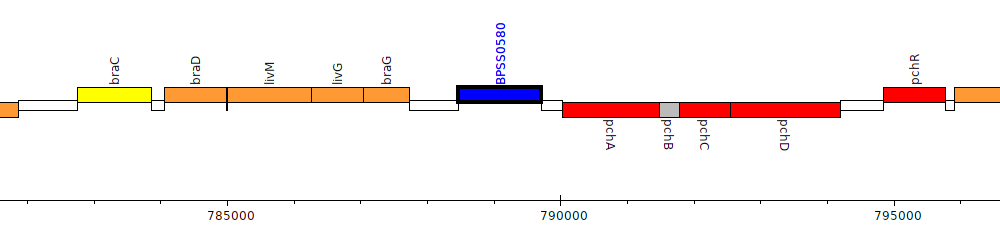
Burkholderia pseudomallei K96243, BPSS0580
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004222 | metalloendopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00933
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:PR00933
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:2.70.70.10 | IPR011055 | Duplicated hybrid motif | 150 | 318 | 6.9E-35 | |
| PRINTS | PR00933 | B-lytic metalloendopeptidase (M23) signature | IPR000841 | Peptidase M23A, B-lytic metalloendopeptidase | 253 | 273 | 5.4E-7 |
| PRINTS | PR00933 | B-lytic metalloendopeptidase (M23) signature | IPR000841 | Peptidase M23A, B-lytic metalloendopeptidase | 204 | 222 | 5.4E-7 |
| Pfam | PF01551 | Peptidase family M23 | IPR016047 | Peptidase M23 | 184 | 275 | 1.6E-14 |
| SMART | SM00287 | IPR003646 | SH3-like domain, bacterial-type | 329 | 401 | 0.0045 | |
| Gene3D | G3DSA:2.30.30.40 | 330 | 398 | 5.5E-6 | |||
| Pfam | PF08239 | Bacterial SH3 domain | IPR003646 | SH3-like domain, bacterial-type | 339 | 397 | 1.4E-7 |
| SUPERFAMILY | SSF51261 | IPR011055 | Duplicated hybrid motif | 167 | 278 | 9.42E-16 | |
| PRINTS | PR00933 | B-lytic metalloendopeptidase (M23) signature | IPR000841 | Peptidase M23A, B-lytic metalloendopeptidase | 157 | 178 | 5.4E-7 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.