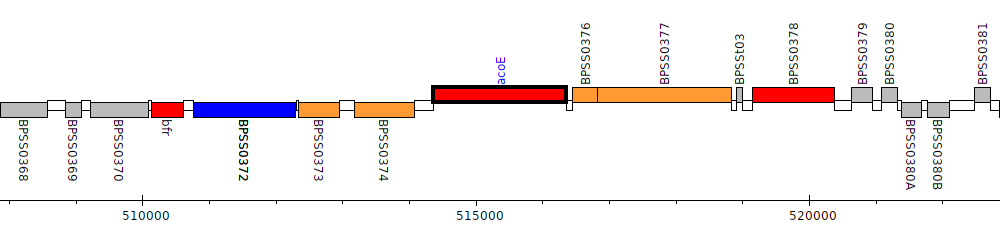
Burkholderia pseudomallei K96243, BPSS0375 (acoE)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003987 | acetate-CoA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02188
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016208 | AMP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02188
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00501
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019427 | acetyl-CoA biosynthetic process from acetate |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02188
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00640 | Propanoate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00680 | Methane metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | chitin degradation to ethanol | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | <i>cis</i>-genanyl-CoA degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00010 | Glycolysis / Gluconeogenesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | colupulone and cohumulone biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-isoleucine biosynthesis V | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00620 | Pyruvate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps00620 | Pyruvate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00630 | Glyoxylate and dicarboxylate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | lupulone and humulone biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00010 | Glycolysis / Gluconeogenesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00720 | Carbon fixation pathways in prokaryotes | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps00640 | Propanoate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | adlupulone and adhumulone biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00680 | Methane metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR02188 | Ac_CoA_lig_AcsA: acetate--CoA ligase | IPR011904 | Acetate-CoA ligase | 28 | 658 | 2.5E-293 |
| SUPERFAMILY | SSF56801 | 17 | 656 | 1.57E-174 | |||
| Pfam | PF13193 | AMP-binding enzyme C-terminal domain | IPR025110 | AMP-binding enzyme, C-terminal domain | 544 | 625 | 4.6E-28 |
| Pfam | PF00501 | AMP-binding enzyme | IPR000873 | AMP-dependent synthetase/ligase | 91 | 535 | 8.6E-95 |
| Gene3D | G3DSA:3.30.300.30 | 531 | 659 | 1.5E-29 | |||
| Hamap | MF_01123 | Acetyl-coenzyme A synthetase [acs]. | IPR011904 | Acetate-CoA ligase | 9 | 660 | 51.609 |
| Pfam | PF16177 | Acetyl-coenzyme A synthetase N-terminus | IPR032387 | Acetyl-coenzyme A synthetase, N-terminal domain | 32 | 87 | 8.6E-21 |
| Gene3D | G3DSA:3.40.50.12780 | IPR042099 | AMP-dependent synthetase-like superfamily | 62 | 528 | 3.0E-101 | |
| ProSitePatterns | PS00455 | Putative AMP-binding domain signature. | IPR020845 | AMP-binding, conserved site | 267 | 278 | - |
| CDD | cd05966 | ACS | 31 | 649 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.