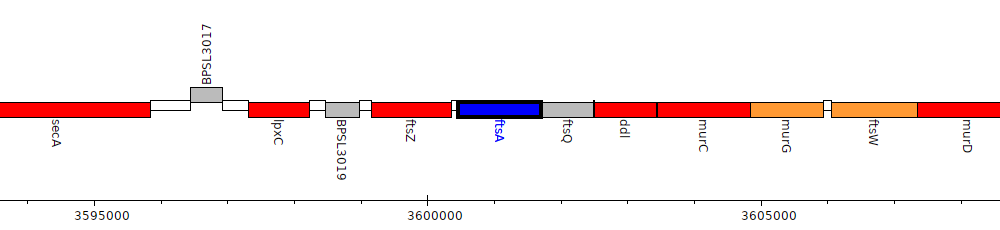
Burkholderia pseudomallei K96243, BPSL3021 (ftsA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0051301 | cell division |
Inferred from Sequence Model
Term mapped from: InterPro:SM00842
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00842
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF14450 | Cell division protein FtsA | 10 | 193 | 2.0E-12 | ||
| PIRSF | PIRSF003101 | IPR020823 | Cell division protein FtsA | 1 | 410 | 3.4E-153 | |
| Gene3D | G3DSA:3.30.420.40 | 12 | 371 | 1.8E-123 | |||
| SUPERFAMILY | SSF53067 | 197 | 375 | 1.09E-49 | |||
| Gene3D | G3DSA:3.30.420.40 | 196 | 351 | 1.8E-123 | |||
| CDD | cd00012 | NBD_sugar-kinase_HSP70_actin | 10 | 230 | 1.85283E-5 | ||
| Pfam | PF02491 | SHS2 domain inserted in FTSA | IPR003494 | SHS2 domain inserted in FtsA | 84 | 161 | 1.2E-32 |
| TIGRFAM | TIGR01174 | ftsA: cell division protein FtsA | IPR020823 | Cell division protein FtsA | 9 | 375 | 1.0E-148 |
| Gene3D | G3DSA:3.90.640.10 | 234 | 299 | 1.8E-123 | |||
| Coils | Coil | 50 | 70 | - | |||
| SMART | SM00842 | IPR003494 | SHS2 domain inserted in FtsA | 9 | 195 | 1.3E-103 | |
| Hamap | MF_02033 | Cell division protein FtsA [ftsA]. | IPR020823 | Cell division protein FtsA | 5 | 410 | 31.559 |
| Gene3D | G3DSA:3.30.1490.110 | 86 | 168 | 1.8E-123 | |||
| SUPERFAMILY | SSF53067 | 7 | 196 | 4.92E-62 | |||
| Pfam | PF14450 | Cell division protein FtsA | 206 | 373 | 4.2E-25 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.