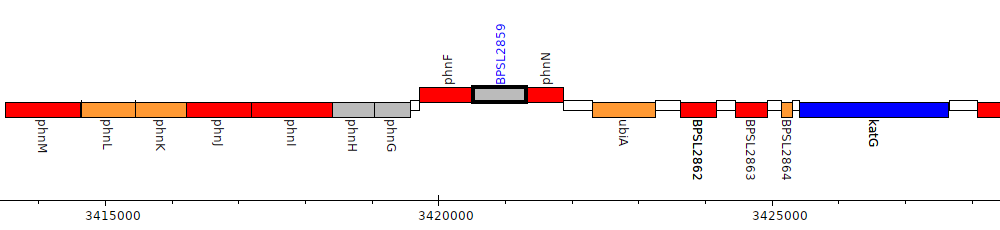
Burkholderia pseudomallei K96243, BPSL2859
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | (aminomethyl)phosphonate degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00030 | Pentose phosphate pathway | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | glyphosate degradation III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR03223 | Phn_opern_protn: putative phosphonate metabolism protein | IPR009389 | Protein of unknown function DUF1045 | 12 | 238 | 3.2E-84 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 245 | 265 | - | ||
| Pfam | PF06299 | Protein of unknown function (DUF1045) | IPR009389 | Protein of unknown function DUF1045 | 71 | 229 | 1.2E-50 |
| PIRSF | PIRSF033328 | IPR009389 | Protein of unknown function DUF1045 | 12 | 247 | 1.0E-44 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.