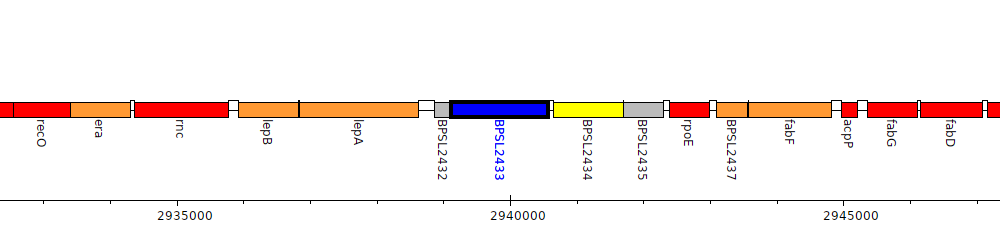
Burkholderia pseudomallei K96243, BPSL2433
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF50156
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004252 | serine-type endopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02037
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004252 | serine-type endopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02037
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02037
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02037
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF50156
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR02037 | degP_htrA_DO: peptidase Do | IPR011782 | Peptidase S1C, Do | 33 | 478 | 6.0E-155 |
| Gene3D | G3DSA:2.30.42.10 | 393 | 485 | 4.5E-15 | |||
| SMART | SM00228 | IPR001478 | PDZ domain | 295 | 362 | 4.7E-14 | |
| CDD | cd00987 | PDZ_serine_protease | 397 | 480 | 2.25828E-14 | ||
| ProSiteProfiles | PS50106 | PDZ domain profile. | IPR001478 | PDZ domain | 283 | 339 | 12.968 |
| PRINTS | PR00834 | HtrA/DegQ protease family signature | IPR001940 | Peptidase S1C | 320 | 332 | 9.5E-53 |
| PRINTS | PR00834 | HtrA/DegQ protease family signature | IPR001940 | Peptidase S1C | 212 | 229 | 9.5E-53 |
| CDD | cd00987 | PDZ_serine_protease | 280 | 369 | 5.48399E-25 | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 378 | 402 | - | ||
| Pfam | PF13180 | PDZ domain | IPR001478 | PDZ domain | 396 | 476 | 1.3E-8 |
| SUPERFAMILY | SSF50156 | IPR036034 | PDZ superfamily | 396 | 483 | 5.78E-14 | |
| SUPERFAMILY | SSF50156 | IPR036034 | PDZ superfamily | 238 | 363 | 5.52E-25 | |
| Pfam | PF13365 | Trypsin-like peptidase domain | 110 | 241 | 1.2E-34 | ||
| Gene3D | G3DSA:2.40.10.120 | 32 | 273 | 1.5E-82 | |||
| Pfam | PF13180 | PDZ domain | IPR001478 | PDZ domain | 295 | 370 | 1.3E-17 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 387 | 402 | - | ||
| PRINTS | PR00834 | HtrA/DegQ protease family signature | IPR001940 | Peptidase S1C | 119 | 131 | 9.5E-53 |
| Gene3D | G3DSA:2.30.42.10 | 275 | 381 | 2.3E-28 | |||
| PRINTS | PR00834 | HtrA/DegQ protease family signature | IPR001940 | Peptidase S1C | 180 | 204 | 9.5E-53 |
| PRINTS | PR00834 | HtrA/DegQ protease family signature | IPR001940 | Peptidase S1C | 234 | 251 | 9.5E-53 |
| SUPERFAMILY | SSF50494 | IPR009003 | Peptidase S1, PA clan | 31 | 272 | 1.2E-67 | |
| PRINTS | PR00834 | HtrA/DegQ protease family signature | IPR001940 | Peptidase S1C | 140 | 160 | 9.5E-53 |
| SMART | SM00228 | IPR001478 | PDZ domain | 395 | 473 | 6.7E-6 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.