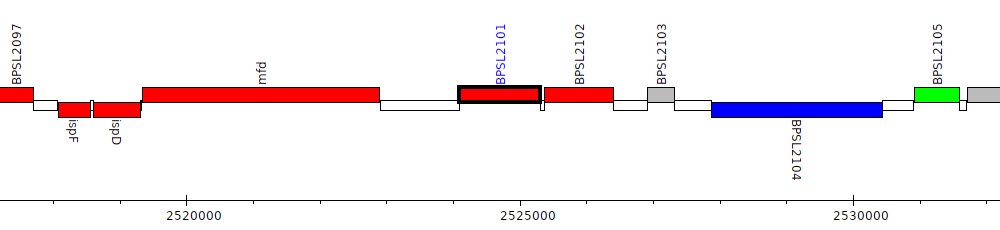
Burkholderia pseudomallei K96243, BPSL2101
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006526 | arginine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01892
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01546
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008777 | acetylornithine deacetylase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01892
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps00330 | Arginine and proline metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01210 | 2-Oxocarboxylic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | L-arginine biosynthesis III (via <i>N</i>-acetyl-L-citrulline) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00220 | Arginine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.30.70.360 | 195 | 312 | 3.2E-103 | |||
| Gene3D | G3DSA:3.40.630.10 | 29 | 388 | 3.2E-103 | |||
| CDD | cd03894 | M20_ArgE | 25 | 400 | 0.0 | ||
| Pfam | PF01546 | Peptidase family M20/M25/M40 | IPR002933 | Peptidase M20 | 88 | 400 | 1.1E-37 |
| SUPERFAMILY | SSF53187 | 23 | 400 | 9.71E-58 | |||
| TIGRFAM | TIGR01892 | AcOrn-deacetyl: acetylornithine deacetylase (ArgE) | IPR010169 | Acetylornithine deacetylase ArgE | 27 | 399 | 6.4E-113 |
| SUPERFAMILY | SSF55031 | IPR036264 | Bacterial exopeptidase dimerisation domain | 195 | 301 | 2.09E-19 | |
| Pfam | PF07687 | Peptidase dimerisation domain | IPR011650 | Peptidase M20, dimerisation domain | 191 | 301 | 7.7E-23 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.