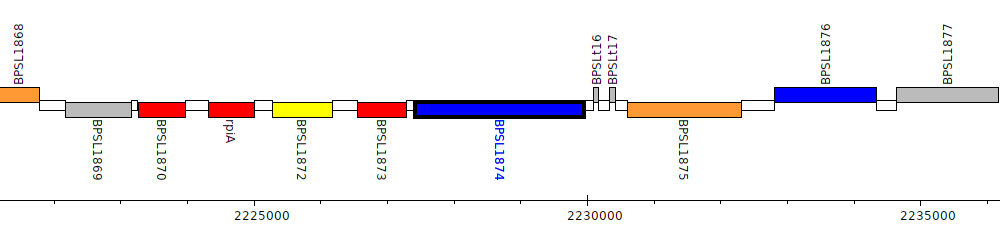
Burkholderia pseudomallei K96243, BPSL1874
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0016070 | RNA metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00358
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004518 | nuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01895
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004518 | nuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01895
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004540 | ribonuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00955
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00955
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00575
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00955
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00575
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004540 | ribonuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00955
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016070 | RNA metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00358
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps03018 | RNA degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:2.40.50.140 | 690 | 787 | 1.5E-19 | |||
| Coils | Coil | 841 | 844 | - | |||
| Pfam | PF17876 | Cold shock domain | IPR040476 | RNase II/RNase R, cold shock domain | 186 | 257 | 2.4E-23 |
| Pfam | PF08206 | Ribonuclease B OB domain | IPR013223 | Ribonuclease B, N-terminal OB domain | 108 | 165 | 1.8E-23 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 780 | 844 | - | ||
| ProSiteProfiles | PS50126 | S1 domain profile. | IPR003029 | S1 domain | 689 | 770 | 14.329 |
| SMART | SM00357 | IPR011129 | Cold shock domain | 107 | 166 | 8.4E-13 | |
| Pfam | PF00773 | RNB domain | IPR001900 | Ribonuclease II/R | 280 | 611 | 7.2E-104 |
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 689 | 769 | 2.34E-20 | |
| Hamap | MF_01895 | Ribonuclease R [rnr]. | IPR011805 | Ribonuclease R | 59 | 770 | 56.325 |
| ProSitePatterns | PS01175 | Ribonuclease II family signature. | IPR022966 | Ribonuclease II/R, conserved site | 579 | 603 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 829 | 844 | - | ||
| Pfam | PF00575 | S1 RNA binding domain | IPR003029 | S1 domain | 686 | 767 | 8.9E-11 |
| CDD | cd04471 | S1_RNase_R | 688 | 770 | 1.40204E-38 | ||
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 88 | 168 | 3.43E-19 | |
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 207 | 687 | 3.43E-142 | |
| TIGRFAM | TIGR00358 | 3_prime_RNase: VacB and RNase II family 3'-5' exoribonucleases | IPR004476 | Ribonuclease II/ribonuclease R | 97 | 770 | 7.8E-213 |
| Gene3D | G3DSA:2.40.50.140 | 93 | 168 | 1.4E-19 | |||
| SMART | SM00316 | IPR022967 | RNA-binding domain, S1 | 687 | 770 | 2.1E-11 | |
| SMART | SM00955 | IPR001900 | Ribonuclease II/R | 280 | 612 | 9.3E-183 | |
| TIGRFAM | TIGR02063 | RNase_R: ribonuclease R | IPR011805 | Ribonuclease R | 41 | 770 | 6.8E-250 |
| SMART | SM00316 | IPR022967 | RNA-binding domain, S1 | 101 | 163 | 11.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.