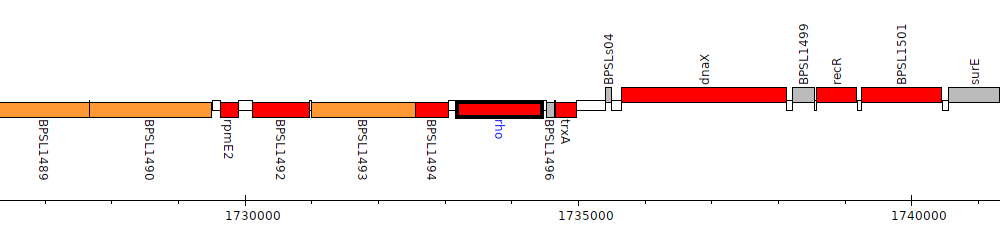
Burkholderia pseudomallei K96243, BPSL1495 (rho)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01884
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01884
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00357
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006353 | DNA-templated transcription, termination |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01884
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008186 | RNA-dependent ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01884
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps03018 | RNA degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00190 | Oxidative phosphorylation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | ATP biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00195 | Photosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00006 | ATP synthase alpha/beta family, nucleotide-binding domain | IPR000194 | ATPase, F1/V1/A1 complex, alpha/beta subunit, nucleotide-binding domain | 160 | 363 | 9.0E-22 |
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 49 | 125 | 1.83E-28 | |
| SUPERFAMILY | SSF68912 | IPR036269 | Rho termination factor, N-terminal domain superfamily | 1 | 46 | 1.96E-12 | |
| Gene3D | G3DSA:1.10.720.10 | 1 | 46 | 3.6E-20 | |||
| TIGRFAM | TIGR00767 | rho: transcription termination factor Rho | IPR004665 | Transcription termination factor Rho | 2 | 417 | 2.0E-209 |
| Hamap | MF_01884 | Transcription termination factor Rho [rho]. | IPR004665 | Transcription termination factor Rho | 1 | 419 | 242.914 |
| CDD | cd01128 | rho_factor | IPR041703 | Transcription termination factor Rho, ATP binding domain | 156 | 404 | 4.51909E-173 |
| Gene3D | G3DSA:2.40.50.140 | 47 | 123 | 3.3E-33 | |||
| SMART | SM00382 | IPR003593 | AAA+ ATPase domain | 170 | 349 | 2.5E-6 | |
| SMART | SM00959 | IPR011112 | Rho termination factor, N-terminal | 5 | 47 | 2.5E-17 | |
| SMART | SM00357 | IPR011129 | Cold shock domain | 52 | 118 | 2.3E-17 | |
| Gene3D | G3DSA:3.40.50.300 | 130 | 417 | 9.2E-128 | |||
| Pfam | PF07498 | Rho termination factor, N-terminal domain | IPR011112 | Rho termination factor, N-terminal | 5 | 47 | 3.6E-16 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 90 | 371 | 7.57E-68 | |
| CDD | cd04459 | Rho_CSD | IPR011113 | Rho termination factor, RNA-binding domain | 51 | 118 | 2.47957E-39 |
| Pfam | PF07497 | Rho termination factor, RNA-binding domain | IPR011113 | Rho termination factor, RNA-binding domain | 52 | 125 | 5.8E-31 |
| ProSiteProfiles | PS51856 | Rho RNA-binding domain profile. | 48 | 123 | 30.616 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.