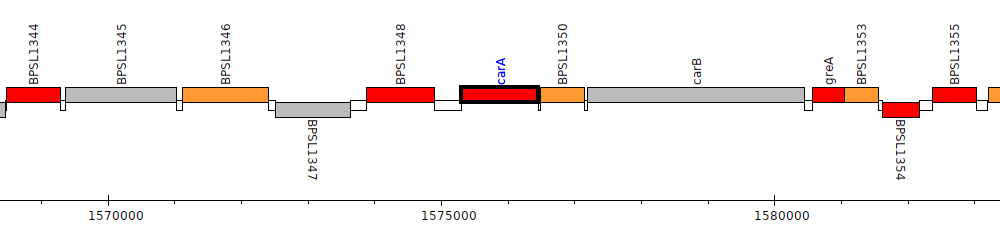
Burkholderia pseudomallei K96243, BPSL1349 (carA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004088 | carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01209
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006541 | glutamine metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01209
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006207 | 'de novo' pyrimidine nucleobase biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01209
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00250 | Alanine, aspartate and glutamate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | L-arginine biosynthesis III (via <i>N</i>-acetyl-L-citrulline) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-arginine biosynthesis IV (archaebacteria) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | UMP biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00250 | Alanine, aspartate and glutamate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00240 | Pyrimidine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps00240 | Pyrimidine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | UMP biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | UMP biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR01368 | CPSaseIIsmall: carbamoyl-phosphate synthase, small subunit | IPR006274 | Carbamoyl-phosphate synthase, small subunit | 8 | 374 | 2.4E-142 |
| Gene3D | G3DSA:3.40.50.880 | IPR029062 | Class I glutamine amidotransferase-like | 154 | 376 | 1.4E-72 | |
| ProSiteProfiles | PS51273 | Glutamine amidotransferase type 1 domain profile. | IPR017926 | Glutamine amidotransferase | 192 | 379 | 27.61 |
| Pfam | PF00117 | Glutamine amidotransferase class-I | IPR017926 | Glutamine amidotransferase | 196 | 370 | 2.2E-45 |
| PRINTS | PR00096 | Glutamine amidotransferase superfamily signature | 348 | 361 | 6.2E-10 | ||
| Pfam | PF00988 | Carbamoyl-phosphate synthase small chain, CPSase domain | IPR002474 | Carbamoyl-phosphate synthase small subunit, N-terminal domain | 8 | 135 | 7.5E-52 |
| PRINTS | PR00099 | Carbamoyl-phosphate synthase protein GATase domain signature | 305 | 316 | 6.5E-39 | ||
| CDD | cd01744 | GATase1_CPSase | IPR035686 | Carbamoyl-phosphate synthase small subunit, GATase1 domain | 193 | 369 | 5.50379E-113 |
| SUPERFAMILY | SSF52021 | IPR036480 | Carbamoyl-phosphate synthase small subunit, N-terminal domain superfamily | 7 | 152 | 2.22E-57 | |
| PRINTS | PR00099 | Carbamoyl-phosphate synthase protein GATase domain signature | 193 | 207 | 6.5E-39 | ||
| PRINTS | PR00096 | Glutamine amidotransferase superfamily signature | 263 | 274 | 6.2E-10 | ||
| PRINTS | PR00099 | Carbamoyl-phosphate synthase protein GATase domain signature | 232 | 246 | 6.5E-39 | ||
| SUPERFAMILY | SSF52317 | IPR029062 | Class I glutamine amidotransferase-like | 158 | 374 | 1.07E-61 | |
| Hamap | MF_01209 | Carbamoyl-phosphate synthase small chain [carA]. | IPR006274 | Carbamoyl-phosphate synthase, small subunit | 6 | 377 | 48.011 |
| PRINTS | PR00096 | Glutamine amidotransferase superfamily signature | 235 | 244 | 6.2E-10 | ||
| PRINTS | PR00099 | Carbamoyl-phosphate synthase protein GATase domain signature | 263 | 279 | 6.5E-39 | ||
| PRINTS | PR00099 | Carbamoyl-phosphate synthase protein GATase domain signature | 280 | 297 | 6.5E-39 | ||
| Gene3D | G3DSA:3.50.30.20 | IPR036480 | Carbamoyl-phosphate synthase small subunit, N-terminal domain superfamily | 6 | 153 | 1.3E-57 | |
| SMART | SM01097 | IPR002474 | Carbamoyl-phosphate synthase small subunit, N-terminal domain | 6 | 136 | 7.7E-90 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.