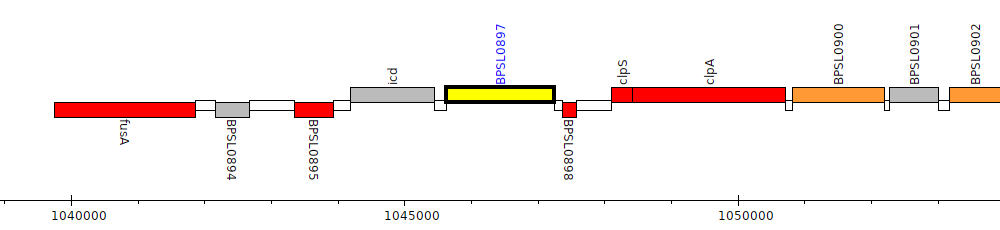
Burkholderia pseudomallei K96243, BPSL0897
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07731
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF07731
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005507 | copper ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00080
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps00860 | Porphyrin and chlorophyll metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF49503 | IPR008972 | Cupredoxin | 422 | 538 | 2.72E-30 | |
| SUPERFAMILY | SSF49503 | IPR008972 | Cupredoxin | 228 | 338 | 2.2E-16 | |
| CDD | cd13881 | CuRO_2_McoC_like | 231 | 378 | 3.97251E-56 | ||
| Gene3D | G3DSA:2.60.40.420 | IPR008972 | Cupredoxin | 221 | 367 | 1.4E-37 | |
| Gene3D | G3DSA:2.60.40.420 | IPR008972 | Cupredoxin | 402 | 538 | 9.5E-40 | |
| Pfam | PF07731 | Multicopper oxidase | IPR011706 | Multicopper oxidase, type 2 | 430 | 538 | 3.9E-28 |
| ProSitePatterns | PS00080 | Multicopper oxidases signature 2. | IPR002355 | Multicopper oxidase, copper-binding site | 521 | 532 | - |
| Gene3D | G3DSA:2.60.40.420 | IPR008972 | Cupredoxin | 64 | 219 | 8.1E-45 | |
| SUPERFAMILY | SSF49503 | IPR008972 | Cupredoxin | 86 | 224 | 1.33E-30 | |
| Pfam | PF07732 | Multicopper oxidase | IPR011707 | Multicopper oxidase, type 3 | 127 | 222 | 1.4E-26 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.