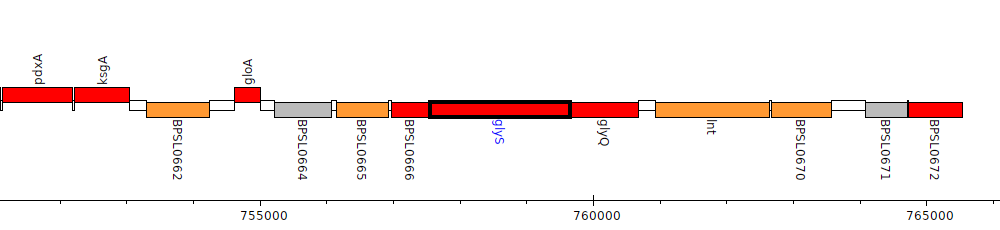
Burkholderia pseudomallei K96243, BPSL0667 (glyS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006426 | glycyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00255
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006420 | arginyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:SM00836
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004814 | arginine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00836
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00255
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004820 | glycine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00255
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00255
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00255
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00970 | Aminoacyl-tRNA biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps00970 | Aminoacyl-tRNA biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR01045 | Glycyl-tRNA synthetase beta subunit signature | IPR015944 | Glycine-tRNA ligase, beta subunit | 255 | 270 | 3.9E-33 |
| PRINTS | PR01045 | Glycyl-tRNA synthetase beta subunit signature | IPR015944 | Glycine-tRNA ligase, beta subunit | 327 | 343 | 3.9E-33 |
| SUPERFAMILY | SSF109604 | 353 | 517 | 5.23E-18 | |||
| PRINTS | PR01045 | Glycyl-tRNA synthetase beta subunit signature | IPR015944 | Glycine-tRNA ligase, beta subunit | 55 | 67 | 3.9E-33 |
| Pfam | PF02092 | Glycyl-tRNA synthetase beta subunit | IPR015944 | Glycine-tRNA ligase, beta subunit | 9 | 559 | 1.4E-197 |
| PRINTS | PR01045 | Glycyl-tRNA synthetase beta subunit signature | IPR015944 | Glycine-tRNA ligase, beta subunit | 10 | 23 | 3.9E-33 |
| SMART | SM00836 | IPR008909 | DALR anticodon binding | 594 | 697 | 3.6E-4 | |
| PRINTS | PR01045 | Glycyl-tRNA synthetase beta subunit signature | IPR015944 | Glycine-tRNA ligase, beta subunit | 403 | 422 | 3.9E-33 |
| Pfam | PF05746 | DALR anticodon binding domain | IPR008909 | DALR anticodon binding | 591 | 689 | 1.1E-9 |
| Hamap | MF_00255 | Glycine--tRNA ligase beta subunit [glyS]. | IPR015944 | Glycine-tRNA ligase, beta subunit | 6 | 697 | 24.039 |
| ProSiteProfiles | PS50861 | Heterodimeric glycyl-transfer RNA synthetases family profile. | IPR006194 | Glycine-tRNA synthetase, heterodimeric | 372 | 662 | 84.612 |
| TIGRFAM | TIGR00211 | glyS: glycine--tRNA ligase, beta subunit | IPR015944 | Glycine-tRNA ligase, beta subunit | 6 | 697 | 2.3E-197 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.