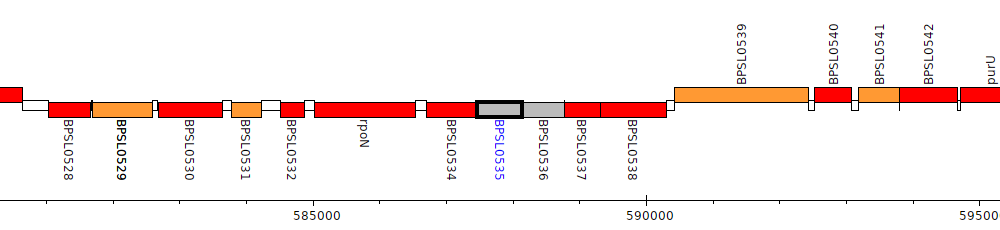
Burkholderia pseudomallei K96243, BPSL0535
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0042597 | periplasmic space |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01914
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0001530 | lipopolysaccharide binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01914
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0015920 | lipopolysaccharide transport |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01914
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:2.60.450.10 | 41 | 198 | 2.0E-45 | |||
| TIGRFAM | TIGR03002 | outer_YhbN_LptA: lipopolysaccharide transport periplasmic protein LptA | IPR014340 | Lipopolysaccharide export system protein LptA | 39 | 195 | 2.6E-45 |
| Pfam | PF03968 | OstA-like protein | IPR005653 | Organic solvent tolerance-like, N-terminal | 49 | 168 | 3.8E-30 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 192 | 221 | - | ||
| Hamap | MF_01914 | Lipopolysaccharide export system protein LptA [lptA]. | IPR014340 | Lipopolysaccharide export system protein LptA | 19 | 195 | 27.046 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.